Dysregulated glucuronic acid metabolism exacerbates hepatocellular carcinoma progression and metastasis through the TGFβ signalling pathway
- PMID: 35979621
- PMCID: PMC9386326
- DOI: 10.1002/ctm2.995
Dysregulated glucuronic acid metabolism exacerbates hepatocellular carcinoma progression and metastasis through the TGFβ signalling pathway
Erratum in
-
Corrigendum to "Dysregulated glucuronic acid metabolism exacerbates hepatocellular carcinoma progression and metastasis through the TGFβ signalling pathway".Clin Transl Med. 2023 Mar;13(3):e1231. doi: 10.1002/ctm2.1231. Clin Transl Med. 2023. PMID: 36941752 Free PMC article. No abstract available.
Abstract
Background: Glucuronic acid metabolism participates in cellular detoxification, extracellular matrix remodeling and cell adhesion and migration. Here, we aimed to explore the crosstalk between dysregulated glucuronic acid metabolism and crucial metastatic signalling in glutathione S-transferase zeta 1 (GSTZ1)-deficient hepatocellular carcinoma (HCC).
Methods: Transwell, HCC xenograft and Gstz1-/- mouse models were used to examine the role of GSTZ1 in HCC metastasis. Non-targeted and targeted metabolomics and global transcriptomic analyses were performed to screen significantly altered metabolic and signalling pathways in GSTZ1 overexpressing hepatoma cells. Further, RNA-binding protein immunoprecipitation, Biotin-RNA pull-down, mRNA decay assays and luciferase reporter assays were used to explore the interaction between RNA and RNA-binding proteins.
Results: GSTZ1 was universally silenced in both human and murine HCC cells, and its deficiency contributed to HCC metastasis in vitro and in vivo. UDP-glucose 6-dehydrogenase (UGDH)-mediated UDP-glucuronic acid (UDP-GlcUA) accumulation promoted hepatoma cell migration upon GSTZ1 loss. UDP-GlcUA stabilized TGFβR1 mRNA by enhancing its binding to polypyrimidine tract binding protein 3, contributing to the activation of TGFβ/Smad signalling. UGDH or TGFβR1 blockade impaired HCC metastasis. In addition, UGDH up-regulation and UDP-GlcUA accumulation correlated with increased metastatic potential and decreased patient survival in GSTZ1-deficient HCC.
Conclusions: GSTZ1 deficiency and subsequent up-regulation of the glucuronic acid metabolic pathway promotes HCC metastasis by increasing the stability of TGFβR1 mRNA and activating TGFβ/Smad signalling. UGDH and a key metabolite, UDP-GlcUA, may serve as prognostic markers. Targeting UGDH might be a promising strategy for HCC therapy.
Keywords: TGFβ/Smad signalling; UDP-GlcUA; UGDH; hepatocellular carcinoma metastasis.
© 2022 The Authors. Clinical and Translational Medicine published by John Wiley & Sons Australia, Ltd on behalf of Shanghai Institute of Clinical Bioinformatics.
Conflict of interest statement
The authors have declared that no conflict of interest exists.
Figures






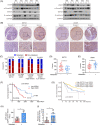

Similar articles
-
GSTZ1 deficiency promotes hepatocellular carcinoma proliferation via activation of the KEAP1/NRF2 pathway.J Exp Clin Cancer Res. 2019 Oct 30;38(1):438. doi: 10.1186/s13046-019-1459-6. J Exp Clin Cancer Res. 2019. PMID: 31666108 Free PMC article.
-
UDP-glucose 6-dehydrogenase lessens sorafenib sensitivity via modulating unfolded protein response.Biochem Biophys Res Commun. 2022 Jul 12;613:207-213. doi: 10.1016/j.bbrc.2022.05.048. Epub 2022 May 14. Biochem Biophys Res Commun. 2022. PMID: 35617808
-
UDP-glucose accelerates SNAI1 mRNA decay and impairs lung cancer metastasis.Nature. 2019 Jul;571(7763):127-131. doi: 10.1038/s41586-019-1340-y. Epub 2019 Jun 26. Nature. 2019. PMID: 31243371
-
UDP-glucose dehydrogenase (UGDH) in clinical oncology and cancer biology.Oncotarget. 2023 Sep 28;14:843-857. doi: 10.18632/oncotarget.28514. Oncotarget. 2023. PMID: 37769033 Free PMC article. Review.
-
UDP-glucose dehydrogenase: structure and function of a potential drug target.Biochem Soc Trans. 2010 Oct;38(5):1378-85. doi: 10.1042/BST0381378. Biochem Soc Trans. 2010. PMID: 20863317 Review.
Cited by
-
Vitamin D alleviates HFD-induced hepatic fibrosis by inhibiting DNMT1 to affect the TGFβ1/Smad3 pathway.iScience. 2024 Oct 28;27(12):111262. doi: 10.1016/j.isci.2024.111262. eCollection 2024 Dec 20. iScience. 2024. PMID: 39713736 Free PMC article.
-
Pan-cancer analysis of UDP-glucose 6-dehydrogenase in human tumors and its function in hepatocellular carcinoma.World J Gastrointest Oncol. 2025 Jul 15;17(7):105981. doi: 10.4251/wjgo.v17.i7.105981. World J Gastrointest Oncol. 2025. PMID: 40697240 Free PMC article.
-
Elevated UDP-glucuronic acid levels mend drug resistance and stress responses via a protease and a transporter in Cryptococcus gattii.Proc Natl Acad Sci U S A. 2025 Apr 29;122(17):e2503960122. doi: 10.1073/pnas.2503960122. Epub 2025 Apr 23. Proc Natl Acad Sci U S A. 2025. PMID: 40267138
-
Serum metabolic profiling of patients with diabetic kidney disease based on gas chromatography-mass spectrometry.Front Mol Biosci. 2025 Mar 17;12:1541440. doi: 10.3389/fmolb.2025.1541440. eCollection 2025. Front Mol Biosci. 2025. PMID: 40166083 Free PMC article.
-
Downregulation of UDP-glucose 6-dehydrogenase predicts adverse outcomes in patients with colorectal cancer and promotes tumorigenesis.Sci Rep. 2025 Jul 1;15(1):21522. doi: 10.1038/s41598-025-07889-4. Sci Rep. 2025. PMID: 40595129 Free PMC article.
References
-
- Fransvea E, Angelotti U, Antonaci S, Giannelli G. Blocking transforming growth factor‐beta up‐regulates E‐cadherin and reduces migration and invasion of hepatocellular carcinoma cells. Hepatology 2008;47:1557‐1566. - PubMed
-
- Lehuédé C, Dupuy F, Rabinovitch R, Jones R, Siegel P. Metabolic plasticity as a determinant of tumor growth and metastasis. Cancer Res 2016;76:5201‐5208. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
