Multi-omics assessment of dilated cardiomyopathy using non-negative matrix factorization
- PMID: 35980883
- PMCID: PMC9387871
- DOI: 10.1371/journal.pone.0272093
Multi-omics assessment of dilated cardiomyopathy using non-negative matrix factorization
Abstract
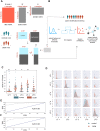
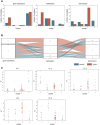
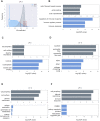
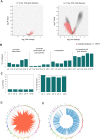
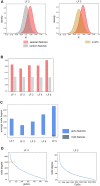
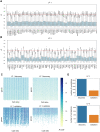
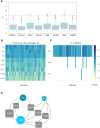
Dilated cardiomyopathy (DCM), a myocardial disease, is heterogeneous and often results in heart failure and sudden cardiac death. Unavailability of cardiac tissue has hindered the comprehensive exploration of gene regulatory networks and nodal players in DCM. In this study, we carried out integrated analysis of transcriptome and methylome data using non-negative matrix factorization from a cohort of DCM patients to uncover underlying latent factors and covarying features between whole-transcriptome and epigenome omics datasets from tissue biopsies of living patients. DNA methylation data from Infinium HM450 and mRNA Illumina sequencing of n = 33 DCM and n = 24 control probands were filtered, analyzed and used as input for matrix factorization using R NMF package. Mann-Whitney U test showed 4 out of 5 latent factors are significantly different between DCM and control probands (P<0.05). Characterization of top 10% features driving each latent factor showed a significant enrichment of biological processes known to be involved in DCM pathogenesis, including immune response (P = 3.97E-21), nucleic acid binding (P = 1.42E-18), extracellular matrix (P = 9.23E-14) and myofibrillar structure (P = 8.46E-12). Correlation network analysis revealed interaction of important sarcomeric genes like Nebulin, Tropomyosin alpha-3 and ERC-protein 2 with CpG methylation of ATPase Phospholipid Transporting 11A0, Solute Carrier Family 12 Member 7 and Leucine Rich Repeat Containing 14B, all with significant P values associated with correlation coefficients >0.7. Using matrix factorization, multi-omics data derived from human tissue samples can be integrated and novel interactions can be identified. Hypothesis generating nature of such analysis could help to better understand the pathophysiology of complex traits such as DCM.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
Epigenome-wide association study for dilated cardiomyopathy in left ventricular heart tissue identifies putative gene sets associated with cardiac pathology and early indicators of cardiac risk.Clin Epigenetics. 2025 Mar 8;17(1):45. doi: 10.1186/s13148-025-01854-8. Clin Epigenetics. 2025. PMID: 40057770 Free PMC article.
-
Multiomics Analysis of Transcriptome, Epigenome, and Genome Uncovers Putative Mechanisms for Dilated Cardiomyopathy.Biomed Res Int. 2021 Mar 29;2021:6653802. doi: 10.1155/2021/6653802. eCollection 2021. Biomed Res Int. 2021. Retraction in: Biomed Res Int. 2024 Mar 20;2024:9898717. doi: 10.1155/2024/9898717. PMID: 33860048 Free PMC article. Retracted.
-
Epigenome-Wide Association Study Identifies Cardiac Gene Patterning and a Novel Class of Biomarkers for Heart Failure.Circulation. 2017 Oct 17;136(16):1528-1544. doi: 10.1161/CIRCULATIONAHA.117.027355. Epub 2017 Aug 24. Circulation. 2017. PMID: 28838933
-
Systems analysis of dilated cardiomyopathy in the next generation sequencing era.Wiley Interdiscip Rev Syst Biol Med. 2018 Jul;10(4):e1419. doi: 10.1002/wsbm.1419. Epub 2018 Feb 27. Wiley Interdiscip Rev Syst Biol Med. 2018. PMID: 29485202 Review.
-
Polymorphisms in genes encoding nonsarcomeric proteins and their role in the pathogenesis of dilated cardiomyopathy.Herz. 2012 Dec;37(8):836-41. doi: 10.1007/s00059-012-3698-6. Herz. 2012. PMID: 23188159 Review.
Cited by
-
An machine learning model to predict quality of life subtypes of disabled stroke survivors.Ann Clin Transl Neurol. 2024 Feb;11(2):404-413. doi: 10.1002/acn3.51960. Epub 2023 Dec 7. Ann Clin Transl Neurol. 2024. PMID: 38059703 Free PMC article.
-
Methylations in dilated cardiomyopathy and heart failure.Front Cardiovasc Med. 2025 Apr 11;12:1559550. doi: 10.3389/fcvm.2025.1559550. eCollection 2025. Front Cardiovasc Med. 2025. PMID: 40290189 Free PMC article. Review.
References
-
- Elliott Perry, Andersson Bert, Arbustini Eloisa, Bilinska Zofia, Cecchi Franco, Charron Philippe, et al.. Classification of the cardiomyopathies: a position statement from the European society of cardiology working group on myocardial and pericardial diseases, European Heart Journal, Volume 29, Issue 2, January 2008, Pages 270–276, doi: 10.1093/eurheartj/ehm342 - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

