Construction of an Epithelial-Mesenchymal Transition-Related Model for Clear Cell Renal Cell Carcinoma Prognosis Prediction
- PMID: 35983409
- PMCID: PMC9381281
- DOI: 10.1155/2022/3780391
Construction of an Epithelial-Mesenchymal Transition-Related Model for Clear Cell Renal Cell Carcinoma Prognosis Prediction
Abstract
Background: A rising amount of data demonstrates that the epithelial-mesenchymal transition (EMT) in clear cell renal cell carcinomas (ccRCC) is connected with the advancement of the cancer. In order to understand the role of EMT in ccRCC, it is critical to integrate molecules involved in EMT into prognosis prediction. The objective of this project was to establish a prognosis prediction model using genes associated with EMT in ccRCC.
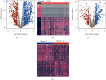
Methods: We acquired the mRNA expression profiles and clinical information about ccRCC from TCGA database. In this study, we measured differentially expressed EMT-related genes (DEEGs) by two comparison groups (tumor versus normal tissues; "stages I-II" versus "stages III-IV" tumor tissues). Based on classification and regression random forest models, we identified the most important DEEGs in predicting prognosis. Afterwards, a risk-score model was created using the identified important DEEGs. The prediction ability of the risk-score model was calculated by the area under the curve (AUC). A nomogram for prognosis prediction was built using the risk-score in combination with clinical factors.
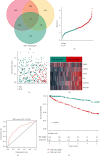
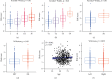
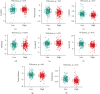
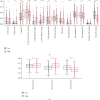
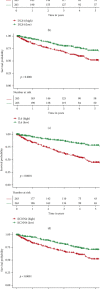
Results: Among the 72 DEEGs, the classification and regression random forest models identified six hub genes (DKK1, DLX4, IL6, KCNN4, RPL22L1, and SPDEF), which exhibited the highest importance values in both models. Through the expression of these six hub genes, a novel risk-score was developed for the prognosis prediction of ccRCC. ROC curves showed the risk-score performed well in both the training (0.749) and testing (0.777) datasets. According to the survival analysis, individuals who were separated into high/low-risk groups had statistically different outcomes in terms of prognosis. Besides, the risk-score model also showed outstanding ability in assessing the progression of ccRCC after treatment. In terms of nomogram, the concordance index (C-index) was 0.79. Additionally, we predicted the differences in response to chemotherapy drugs among patients from low- and high-risk groups.
Conclusion: Gene signatures related to EMT could be useful in predicting ccRCC prognosis.
Copyright © 2022 Shimiao Zhu et al.
Conflict of interest statement
The authors state that they have no conflicts of interest.
Figures

Similar articles
-
Prognostic value of epithelial-mesenchymal transition markers in clear cell renal cell carcinoma.Aging (Albany NY). 2020 Jan 8;12(1):866-883. doi: 10.18632/aging.102660. Epub 2020 Jan 8. Aging (Albany NY). 2020. PMID: 31915310 Free PMC article.
-
Molecular Characterization of Clear Cell Renal Cell Carcinoma Reveals Prognostic Significance of Epithelial-mesenchymal Transition Gene Expression Signature.Eur Urol Oncol. 2022 Feb;5(1):92-99. doi: 10.1016/j.euo.2021.10.007. Epub 2021 Nov 26. Eur Urol Oncol. 2022. PMID: 34840106
-
Identification of an independent immune-genes prognostic index for renal cell carcinoma.BMC Cancer. 2021 Jun 29;21(1):746. doi: 10.1186/s12885-021-08367-6. BMC Cancer. 2021. PMID: 34187413 Free PMC article.
-
The Correlation Between the Immune and Epithelial-Mesenchymal Transition Signatures Suggests Potential Therapeutic Targets and Prognosis Prediction Approaches in Kidney Cancer.Sci Rep. 2018 Apr 26;8(1):6570. doi: 10.1038/s41598-018-25002-w. Sci Rep. 2018. PMID: 29700419 Free PMC article.
-
Large-scale transcriptome profiles reveal robust 20-signatures metabolic prediction models and novel role of G6PC in clear cell renal cell carcinoma.J Cell Mol Med. 2020 Aug;24(16):9012-9027. doi: 10.1111/jcmm.15536. Epub 2020 Jun 21. J Cell Mol Med. 2020. PMID: 32567187 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

