Thymocyte regulatory variant alters transcription factor binding and protects from type 1 diabetes in infants
- PMID: 35986039
- PMCID: PMC9391468
- DOI: 10.1038/s41598-022-18296-4
Thymocyte regulatory variant alters transcription factor binding and protects from type 1 diabetes in infants
Abstract
We recently mapped a genetic susceptibility locus on chromosome 6q22.33 for type 1 diabetes (T1D) diagnosed below the age of 7 years between the PTPRK and thymocyte-selection-associated (THEMIS) genes. As the thymus plays a central role in shaping the T cell repertoire, we aimed to identify the most likely causal genetic factors behind this association using thymocyte genomic data. In four thymocyte populations, we identified 253 DNA sequence motifs underlying histone modifications. The G insertion allele of rs138300818, associated with protection from diabetes, created thymocyte motifs for multiple histone modifications and thymocyte types. In a parallel approach to identifying variants that alter transcription factor binding motifs, the same variant disrupted a predicted motif for Rfx7, which is abundantly expressed in the thymus. Chromatin state and RNA sequencing data suggested strong transcription overlapping rs138300818 in fetal thymus, while expression quantitative trait locus and chromatin conformation data associate the insertion with lower THEMIS expression. Extending the analysis to other T1D loci further highlighted rs66733041 affecting the GATA3 transcription factor binding in the AFF3 locus. Taken together, our results support a role for thymic THEMIS gene expression and the rs138300818 variant in promoting the development of early-onset T1D.
© 2022. The Author(s).
Conflict of interest statement
J.R.J.I. is currently an employee of Exploristics UK. A.J.C. is currently an employee of GSK UK. Other authors report no competing interests.
Figures



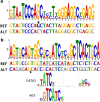
References
-
- Inshaw, J. R. J., Cutler, A. J., Burren, O. S., Stefana, M. I. & Todd, J. A. Approaches and advances in the genetic causes of autoimmune disease and their implications. Nat. Immunol. (2018). - PubMed
-
- Crouch, D. J. et al. Enhanced genetic analysis of type 1 diabetes by selecting variants on both effect size and significance, and by integration with autoimmune thyroid disease. bioRxiv 2021.02.05.429962 (2021).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

