Hypoxia-induced lncRNA STEAP3-AS1 activates Wnt/β-catenin signaling to promote colorectal cancer progression by preventing m6A-mediated degradation of STEAP3 mRNA
- PMID: 35986274
- PMCID: PMC9392287
- DOI: 10.1186/s12943-022-01638-1
Hypoxia-induced lncRNA STEAP3-AS1 activates Wnt/β-catenin signaling to promote colorectal cancer progression by preventing m6A-mediated degradation of STEAP3 mRNA
Abstract
Background: Hypoxia, a typical hallmark of solid tumors, exhibits an essential role in the progression of colorectal cancer (CRC), in which the dysregulation of long non-coding RNAs (lncRNAs) is frequently observed. However, the underlying mechanisms are not clearly defined.
Methods: The TCGA database was analyzed to identify differential lncRNA expression involved in hypoxia-induced CRC progression. qRT-PCR was conducted to validate the upregulation of lncRNA STEAP3-AS1 in CRC cell lines and tumor-bearing mouse and zebrafish models under hypoxia. ChIP-qRT-PCR was used to detect the transcriptional activation of STEAP3-AS1 mediated by HIF-1α. RNA-seq, fluorescent in situ hybridization, RNA pulldown, RNA immunoprecipitation, co-immunoprecipitation, immunofluorescence and immunoblot experiments were used to ascertain the involved mechanisms. Functional assays were performed in both in vitro and in vivo models to investigate the regulatory role of STEAP3-AS1/STEAP3/Wnt/β-catenin axis in CRC proliferation and metastasis.
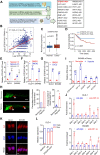
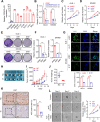
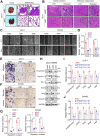
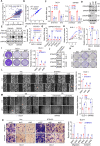
Results: Here, we identified a hypoxia-induced antisense lncRNA STEAP3-AS1 that was highly expressed in clinical CRC tissues and positively correlated with poor prognosis of CRC patients. Upregulation of lncRNA STEAP3-AS1, which was induced by HIF-1α-mediated transcriptional activation, facilitated the proliferation and metastasis of CRC cells both in vitro and in vivo. Mechanistically, STEAP3-AS1 interacted competitively with the YTH domain-containing family protein 2 (YTHDF2), a N6-methyladenosine (m6A) reader, leading to the disassociation of YTHDF2 with STEAP3 mRNA. This effect protected STEAP3 mRNA from m6A-mediated degradation, enabling the high expression of STEAP3 protein and subsequent production of cellular ferrous iron (Fe2+). Increased Fe2+ levels elevated Ser 9 phosphorylation of glycogen synthase kinase 3 beta (GSK3β) and inhibited its kinase activity, thus releasing β-catenin for nuclear translocation and subsequent activation of Wnt signaling to support CRC progression.
Conclusions: Taken together, our study highlights the mechanisms of lncRNA STEAP3-AS1 in facilitating CRC progression involving the STEAP3-AS1/STEAP3/Wnt/β-catenin axis, which may provide novel diagnostic biomarkers or therapeutic targets to benefit CRC treatment. Hypoxia-induced HIF-1α transcriptionally upregulates the expression of lncRNA STEAP3-AS1, which interacts competitively with YTHDF2, thus upregulating mRNA stability of STEAP3 and consequent STEAP3 protein expression. The enhanced STEAP3 expression results in production of cellular ferrous iron (Fe2+), which induces the Ser 9 phosphorylation and inactivation of GSK3β, releasing β-catenin for nuclear translocation and contributing to subsequent activation of Wnt signaling to promote CRC progression.
Keywords: Colorectal cancer; Hypoxia; LncRNA STEAPS-AS1; STEAP3; Wnt/β-catenin; YTHDF2; m6A modification.
© 2022. The Author(s).
Conflict of interest statement
The authors declared no potential competing interests.
Figures
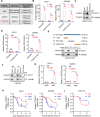

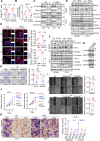
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

