Circular RNA circFIRRE drives osteosarcoma progression and metastasis through tumorigenic-angiogenic coupling
- PMID: 35986280
- PMCID: PMC9389772
- DOI: 10.1186/s12943-022-01624-7
Circular RNA circFIRRE drives osteosarcoma progression and metastasis through tumorigenic-angiogenic coupling
Abstract
Background: Disappointing clinical efficacy of standard treatment has been proven in refractory metastatic osteosarcoma, and the emerging anti-angiogenic regimens are still in the infantile stage. Thus, there is an urgent need to develop novel therapeutic approach for osteosarcoma lung metastasis.
Methods: circFIRRE was selected from RNA-sequencing of 4 matched osteosarcoma and adjacent samples. The expression of circFIRRE was verified in clinical osteosarcoma samples and cell lines via quantitative real-time polymerase chain reaction (RT-qPCR). The effect of circFIRRE was investigated in cell lines in vitro models, ex vivo models and in vivo xenograft tumor models, including proliferation, invasion, migration, metastasis and angiogenesis. Signaling regulatory mechanism was evaluated by RT-qPCR, Western blot, RNA pull-down and dual-luciferase reporter assays.
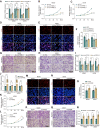
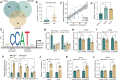
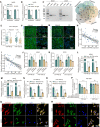
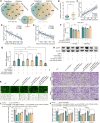
Results: In this article, a novel circular RNA, circFIRRE (hsa_circ_0001944) was screened out and identified from RNA-sequencing, and was upregulated in both osteosarcoma cell lines and tissues. Clinically, aberrantly upregulated circFIRRE portended higher metastatic risk and worse prognosis in osteosarcoma patients. Functionally, in vitro, ex vivo and in vivo experiments demonstrated that circFIRRE could drive primary osteosarcoma progression and lung metastasis by inducing both tumor cells and blood vessels, we call as "tumorigenic-angiogenic coupling". Mechanistically, upregulated circFIRRE was induced by transcription factor YY1, and partially boosted the mRNA and protein level of LUZP1 by sponging miR-486-3p and miR-1225-5p.
Conclusions: We identified circFIRRE as a master regulator in the tumorigenesis and angiogenesis of osteosarcoma, which could be purposed as a novel prognostic biomarker and therapeutic target for refractory osteosarcoma.
Keywords: Angiogenesis; LUZP1; Osteosarcoma; YY1; circFIRRE; miR-1225-5p; miR-486-3p.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



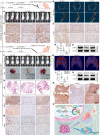
References
-
- Bielack S, Kempf-Bielack B, Delling G, Exner G, Flege S, Helmke K, Kotz R, Salzer-Kuntschik M, Werner M, Winkelmann W, et al. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2002;20:776–90. - PubMed
-
- Zambo I, Veselý K. WHO classification of tumours of soft tissue and bone 2013: the main changes compared to the 3rd edition. Cesk Patol. 2014;50:64–70. - PubMed
-
- Kreuter M, Bieker R, Bielack S, Auras T, Buerger H, Gosheger G, Jurgens H, Berdel W, Mesters R. Prognostic relevance of increased angiogenesis in osteosarcoma. Clinical cancer research : an official journal of the American Association for Cancer Research. 2004;10:8531–7. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

