Selective and comparative genome architecture of Asian cultivated rice (Oryza sativa L.) attributed to domestication and modern breeding
- PMID: 35988902
- PMCID: PMC9788959
- DOI: 10.1016/j.jare.2022.08.004
Selective and comparative genome architecture of Asian cultivated rice (Oryza sativa L.) attributed to domestication and modern breeding
Abstract
Introduction: Rice, Oryza sativa L. (Os), is one of the oldest domesticated cereals that has also gone through extensive improvement in modern breeding.
Objectives: How rice was domesticated and impacted by modern breeding.
Methods: We performed comprehensive analyses of genomic sequences of 504 accessions of Os and 456 accessions of O. rufipogon/O. nivara (Or).
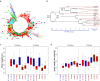
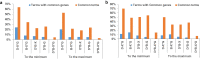
Results: The natural selection on Or before domestication and the natural and artificial selection during domestication together shaped the well-differentiated genomes of two subspecies, geng(j) (japonica) and xian(i) (indica), while breeding has made apparent genomic imprints between landrace and modern varieties of each subspecies, and also between primary modern and advanced modern varieties of xian(i). Selection during domestication and breeding left genome-wide selective signals covering ∼ 22.8 % and ∼ 8.6 % of the Os genome, significantly reduced within-population genomic diversity by ∼ 22 % in xian(i) and ∼ 53 % in geng(j) plus more pronounced subspecific differentiation. Only ∼ 10 % reduction in the total genomic diversity was observed between the Os and Or populations, indicating domestication did not suffer severe genetic bottleneck.
Conclusion: Our results revealed clear differentiation of the Or accessions into three large populations, two of which correspond to the well-differentiated Os subspecies, geng(j) and xian(i). Improved productivity and common changes in the same suit of adaptive traits in xian(i) and geng(j) during domestication and breeding resulted apparently from compensatory and convergent selections for different genes/alleles acting in the common KEGG terms and/or same gene families, and thus maintaining or even increasing the within population diversity and subspecific differentiation of Os, while more genes/alleles of novel function were selected during domestication than modern breeding. Our results supported the multiple independent domestication of Os in Asia and suggest the more efficient utilization of the rich diversity within Os by exploiting inter-subspecific and among population diversity in future rice improvement.
Keywords: Domestication; Modern breeding; Novel functional variation selection; Oryza sativa; Standing functional variation selection.
Copyright © 2022. Production and hosting by Elsevier B.V.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures





References
-
- Zhang D, Zhang H, Wang M, Sun J, Qi Y, Wang F, et al. Genetic Structure and Differentiation of Oryza Sativa L. In China Revealed by Microsatellites. Theor Appl Genet. 2009;119(6):1105–1117. - PubMed
-
- Zhang LB, Zhu Q, Wu ZQ, Ross‐Ibarra J, Gaut BS, Ge S, et al. Selection on Grain Shattering Genes and Rates of Rice Domestication. New Phytol. 2009;184(3):708–720. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

