Inference of putative cell-type-specific imprinted regulatory elements and genes during human neuronal differentiation
- PMID: 35994039
- PMCID: PMC9851749
- DOI: 10.1093/hmg/ddac207
Inference of putative cell-type-specific imprinted regulatory elements and genes during human neuronal differentiation
Abstract
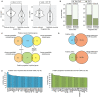
Genomic imprinting results in gene expression bias caused by parental chromosome of origin and occurs in genes with important roles during human brain development. However, the cell-type and temporal specificity of imprinting during human neurogenesis is generally unknown. By detecting within-donor allelic biases in chromatin accessibility and gene expression that are unrelated to cross-donor genotype, we inferred imprinting in both primary human neural progenitor cells and their differentiated neuronal progeny from up to 85 donors. We identified 43/20 putatively imprinted regulatory elements (IREs) in neurons/progenitors, and 133/79 putatively imprinted genes in neurons/progenitors. Although 10 IREs and 42 genes were shared between neurons and progenitors, most putative imprinting was only detected within specific cell types. In addition to well-known imprinted genes and their promoters, we inferred novel putative IREs and imprinted genes. Consistent with both DNA methylation-based and H3K27me3-based regulation of imprinted expression, some putative IREs also overlapped with differentially methylated or histone-marked regions. Finally, we identified a progenitor-specific putatively imprinted gene overlapping with copy number variation that is associated with uniparental disomy-like phenotypes. Our results can therefore be useful in interpreting the function of variants identified in future parent-of-origin association studies.
© The Author(s) 2022. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures




Similar articles
-
Integration of CTCF loops, methylome, and transcriptome in differentiating LUHMES as a model for imprinting dynamics of the 15q11-q13 locus in human neurons.Hum Mol Genet. 2024 Sep 19;33(19):1711-1725. doi: 10.1093/hmg/ddae111. Hum Mol Genet. 2024. PMID: 39045627 Free PMC article.
-
PIK3R1 and G0S2 are human placenta-specific imprinted genes associated with germline-inherited maternal DNA methylation.Epigenetics. 2025 Dec;20(1):2523191. doi: 10.1080/15592294.2025.2523191. Epub 2025 Jun 26. Epigenetics. 2025. PMID: 40568952 Free PMC article.
-
Diagnostic test accuracy and cost-effectiveness of tests for codeletion of chromosomal arms 1p and 19q in people with glioma.Cochrane Database Syst Rev. 2022 Mar 2;3(3):CD013387. doi: 10.1002/14651858.CD013387.pub2. Cochrane Database Syst Rev. 2022. PMID: 35233774 Free PMC article.
-
Comprehensive analysis of imprinted genes in citrus endosperm and their contributions to seed development.Plant J. 2025 Jun;122(6):e70290. doi: 10.1111/tpj.70290. Plant J. 2025. PMID: 40560793
-
Incentives for preventing smoking in children and adolescents.Cochrane Database Syst Rev. 2017 Jun 6;6(6):CD008645. doi: 10.1002/14651858.CD008645.pub3. Cochrane Database Syst Rev. 2017. PMID: 28585288 Free PMC article.
Cited by
-
Nicotinamide Mononucleotide (NMN) Works in Type 2 Diabetes through Unexpected Effects in Adipose Tissue, Not by Mitochondrial Biogenesis.Int J Mol Sci. 2024 Feb 23;25(5):2594. doi: 10.3390/ijms25052594. Int J Mol Sci. 2024. PMID: 38473844 Free PMC article.
References
-
- Bonthuis, P.J., Huang, W.-C., Stacher Hörndli, C.N., Ferris, E., Cheng, T. and Gregg, C. (2015) Noncanonical genomic imprinting effects in offspring. Cell Rep., 12, 979–991. - PubMed
-
- Zink, F., Magnusdottir, D.N., Magnusson, O.T., Walker, N.J., Morris, T.J., Sigurdsson, A., Halldorsson, G.H., Gudjonsson, S.A., Melsted, P., Ingimundardottir, H. et al. (2018) Insights into imprinting from parent-of-origin phased methylomes and transcriptomes. Nat. Genet., 50, 1542–1552. - PubMed
-
- Nakabayashi, K., Trujillo, A.M., Tayama, C., Camprubi, C., Yoshida, W., Lapunzina, P., Sanchez, A., Soejima, H., Aburatani, H., Nagae, G. et al. (2011) Methylation screening of reciprocal genome-wide UPDs identifies novel human-specific imprinted genes. Hum. Mol. Genet., 20, 3188–3197. - PubMed

