Inferring tumor-specific cancer dependencies through integrating ex vivo drug response assays and drug-protein profiling
- PMID: 35994503
- PMCID: PMC9436053
- DOI: 10.1371/journal.pcbi.1010438
Inferring tumor-specific cancer dependencies through integrating ex vivo drug response assays and drug-protein profiling
Abstract
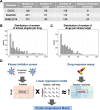
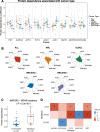
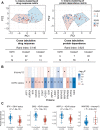
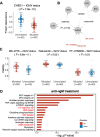
The development of cancer therapies may be improved by the discovery of tumor-specific molecular dependencies. The requisite tools include genetic and chemical perturbations, each with its strengths and limitations. Chemical perturbations can be readily applied to primary cancer samples at large scale, but mechanistic understanding of hits and further pharmaceutical development is often complicated by the fact that a chemical compound has affinities to multiple proteins. To computationally infer specific molecular dependencies of individual cancers from their ex vivo drug sensitivity profiles, we developed a mathematical model that deconvolutes these data using measurements of protein-drug affinity profiles. Through integrating a drug-kinase profiling dataset and several drug response datasets, our method, DepInfeR, correctly identified known protein kinase dependencies, including the EGFR dependence of HER2+ breast cancer cell lines, the FLT3 dependence of acute myeloid leukemia (AML) with FLT3-ITD mutations and the differential dependencies on the B-cell receptor pathway in the two major subtypes of chronic lymphocytic leukemia (CLL). Furthermore, our method uncovered new subgroup-specific dependencies, including a previously unreported dependence of high-risk CLL on Checkpoint kinase 1 (CHEK1). The method also produced a detailed map of the kinase dependencies in a heterogeneous set of 117 CLL samples. The ability to deconvolute polypharmacological phenotypes into underlying causal molecular dependencies should increase the utility of high-throughput drug response assays for functional precision oncology.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
Bayesian inference for identifying tumour-specific cancer dependencies through integration of ex-vivo drug response assays and drug-protein profiling.BMC Bioinformatics. 2024 Mar 8;25(1):104. doi: 10.1186/s12859-024-05682-0. BMC Bioinformatics. 2024. PMID: 38459430 Free PMC article.
-
Survey of ex vivo drug combination effects in chronic lymphocytic leukemia reveals synergistic drug effects and genetic dependencies.Leukemia. 2020 Nov;34(11):2934-2950. doi: 10.1038/s41375-020-0846-5. Epub 2020 May 13. Leukemia. 2020. PMID: 32404973 Free PMC article.
-
LT-171-861, a novel FLT3 inhibitor, shows excellent preclinical efficacy for the treatment of FLT3 mutant acute myeloid leukemia.Theranostics. 2021 Jan 1;11(1):93-106. doi: 10.7150/thno.46593. eCollection 2021. Theranostics. 2021. PMID: 33391463 Free PMC article.
-
Pharmacological impact of FLT3 mutations on receptor activity and responsiveness to tyrosine kinase inhibitors.Biochem Pharmacol. 2021 Jan;183:114348. doi: 10.1016/j.bcp.2020.114348. Epub 2020 Nov 23. Biochem Pharmacol. 2021. PMID: 33242449 Review.
-
FLT3 Inhibitors in Acute Myeloid Leukemia: Current Status and Future Directions.Mol Cancer Ther. 2017 Jun;16(6):991-1001. doi: 10.1158/1535-7163.MCT-16-0876. Mol Cancer Ther. 2017. PMID: 28576946 Free PMC article. Review.
Cited by
-
Bayesian inference for identifying tumour-specific cancer dependencies through integration of ex-vivo drug response assays and drug-protein profiling.BMC Bioinformatics. 2024 Mar 8;25(1):104. doi: 10.1186/s12859-024-05682-0. BMC Bioinformatics. 2024. PMID: 38459430 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

