Polycomb-mediated repression of paternal chromosomes maintains haploid dosage in diploid embryos of Marchantia
- PMID: 35996955
- PMCID: PMC9402228
- DOI: 10.7554/eLife.79258
Polycomb-mediated repression of paternal chromosomes maintains haploid dosage in diploid embryos of Marchantia
Abstract
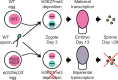
Complex mechanisms regulate gene dosage throughout eukaryotic life cycles. Mechanisms controlling gene dosage have been extensively studied in animals, however it is unknown how generalizable these mechanisms are to diverse eukaryotes. Here, we use the haploid plant Marchantia polymorpha to assess gene dosage control in its short-lived diploid embryo. We show that throughout embryogenesis, paternal chromosomes are repressed resulting in functional haploidy. The paternal genome is targeted for genomic imprinting by the Polycomb mark H3K27me3 starting at fertilization, rendering the maternal genome in control of embryogenesis. Maintaining haploid gene dosage by this new form of imprinting is essential for embryonic development. Our findings illustrate how haploid-dominant species can regulate gene dosage through paternal chromosome inactivation and initiates the exploration of the link between life cycle history and gene dosage in a broader range of organisms.
Keywords: Marchantia polymorpha; Polycomb; chromosomes; embryogenesis; epigenetics; gene dosage; gene expression; genetics; genomic imprinting; genomics.
Plain language summary
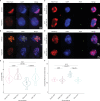
The reproductive cells of organisms that reproduce sexually – the egg and the sperm – each contain one copy of the organism’s genome. An embryo forms upon fertilization of an egg by a sperm cell. This embryo contains two copies of the genome, one from each parent. Under most circumstances, it does not matter which parent a gene copy came from: both gene copies are expressed. However, in some species genes coming from only one of the parents are switched on. This unusual mode of gene expression is called genomic imprinting. The best-known example of this occurs in female mammals, which repress the genes on the paternal X chromosome. Genomic imprinting also exists in flowering plants. Both mammals and flowering plants evolved tissues that channel nutrients from the mother to the embryo during development; the placenta and the endosperm, respectively. Genomic imprinting had, until now, only been described in these two types of organisms. It was unknown whether imprinting also happens in other organisms, and specifically those in which embryos develop inside the mother but without the help of a placenta or endosperm. Here Montgomery et al. addressed this question by studying the liverwort, Marchantia polymorpha, a moss-like plant. Initial experiments showed that cells in the liverwort embryo mostly expressed the genes coming from the egg, and not the sperm. All the genetic material coming from the sperm had a molecular marker or tag called H3K27me3. This mark, which also appears on the paternal X chromosome in female mammals, switches off the genes it tags. M. polymorpha embryos thus suppress gene expression from all of the genetic material from the father, relying only on maternal genetic material for development. When Montgomery et al. deleted the maternal genes necessary for making the H3K27me3 mark, the paternal genes switched on, and this led to the death of the embryos. The survival of M. polymorpha embryos therefore depended on keeping only one set of genes active. Taken together these experiments indicate that genomic imprinting evolved about 480 million years ago, about 320 million years earlier than previously thought, in organisms for which embryo development depended only on one parent. This means there are likely many more organisms that control gene expression in this way, opening up opportunities for further research. Understanding imprinting in more detail will also shed light on how sexual reproduction evolved.
© 2022, Montgomery et al.
Conflict of interest statement
SM, TH, NW, EA, SA, MS, CL, FB No competing interests declared
Figures




Similar articles
-
Paternal imprinting in Marchantia polymorpha.New Phytol. 2024 Feb;241(3):1000-1006. doi: 10.1111/nph.19377. Epub 2023 Nov 7. New Phytol. 2024. PMID: 37936346 Review.
-
Gamete expression of TALE class HD genes activates the diploid sporophyte program in Marchantia polymorpha.Elife. 2021 Sep 17;10:e57088. doi: 10.7554/eLife.57088. Elife. 2021. PMID: 34533136 Free PMC article.
-
The conflict theory of genomic imprinting: how much can be explained?Curr Top Dev Biol. 1998;40:255-93. doi: 10.1016/s0070-2153(08)60369-5. Curr Top Dev Biol. 1998. PMID: 9673853 Review.
-
Identification of the sex-determining factor in the liverwort Marchantia polymorpha reveals unique evolution of sex chromosomes in a haploid system.Curr Biol. 2021 Dec 20;31(24):5522-5532.e7. doi: 10.1016/j.cub.2021.10.023. Epub 2021 Nov 3. Curr Biol. 2021. PMID: 34735792 Free PMC article.
-
Genomic imprinting in plants: what makes the functions of paternal and maternal genes different in endosperm formation?Adv Genet. 2014;86:1-25. doi: 10.1016/B978-0-12-800222-3.00001-2. Adv Genet. 2014. PMID: 25172343 Review.
Cited by
-
Imprinting as Basis for Complex Evolutionary Novelties in Eutherians.Biology (Basel). 2024 Aug 31;13(9):682. doi: 10.3390/biology13090682. Biology (Basel). 2024. PMID: 39336109 Free PMC article. Review.
-
Population Genomics of the Facultatively Sexual Liverwort Marchantia polymorpha.Genome Biol Evol. 2023 Nov 1;15(11):evad196. doi: 10.1093/gbe/evad196. Genome Biol Evol. 2023. PMID: 37883717 Free PMC article.
-
Simultaneous Visualization of Protein and Genomic Regions in Plant Nuclei.Methods Mol Biol. 2025;2873:205-222. doi: 10.1007/978-1-0716-4228-3_12. Methods Mol Biol. 2025. PMID: 39576604
-
Reactivation of the tRNASer/tRNATyr gene cluster in Arabidopsis thaliana root tips.Plant Cell. 2025 Jul 1;37(7):koaf137. doi: 10.1093/plcell/koaf137. Plant Cell. 2025. PMID: 40479509 Free PMC article.
-
Interactions between U and V sex chromosomes during the life cycle of Ectocarpus.Development. 2024 Oct 15;151(20):dev202677. doi: 10.1242/dev.202677. Epub 2024 Apr 11. Development. 2024. PMID: 38512707 Free PMC article.
References
-
- Anderson SN, Johnson CS, Chesnut J, Jones DS, Khanday I, Woodhouse M, Li C, Conrad LJ, Russell SD, Sundaresan V. The zygotic transition is initiated in unicellular plant zygotes with asymmetric activation of parental genomes. Developmental Cell. 2017;43:349–358. doi: 10.1016/j.devcel.2017.10.005. - DOI - PubMed
-
- Bachtrog D, Mank JE, Peichel CL, Kirkpatrick M, Otto SP, Ashman TL, Hahn MW, Kitano J, Mayrose I, Ming R, Perrin N, Ross L, Valenzuela N, Vamosi JC, Tree of Sex Consortium Sex determination: why so many ways of doing it? PLOS Biology. 2014;12:e1001899. doi: 10.1371/journal.pbio.1001899. - DOI - PMC - PubMed
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

