Phenotype Harmonization in the GLIDE2 Oral Health Genomics Consortium
- PMID: 36000800
- PMCID: PMC9516613
- DOI: 10.1177/00220345221109775
Phenotype Harmonization in the GLIDE2 Oral Health Genomics Consortium
Abstract
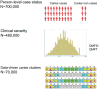
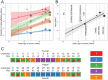
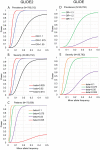
Genetic risk factors play important roles in the etiology of oral, dental, and craniofacial diseases. Identifying the relevant risk loci and understanding their molecular biology could highlight new prevention and management avenues. Our current understanding of oral health genomics suggests that dental caries and periodontitis are polygenic diseases, and very large sample sizes and informative phenotypic measures are required to discover signals and adequately map associations across the human genome. In this article, we introduce the second wave of the Gene-Lifestyle Interactions and Dental Endpoints consortium (GLIDE2) and discuss relevant data analytics challenges, opportunities, and applications. In this phase, the consortium comprises a diverse, multiethnic sample of over 700,000 participants from 21 studies contributing clinical data on dental caries experience and periodontitis. We outline the methodological challenges of combining data from heterogeneous populations, as well as the data reduction problem in resolving detailed clinical examination records into tractable phenotypes, and describe a strategy that addresses this. Specifically, we propose a 3-tiered phenotyping approach aimed at leveraging both the large sample size in the consortium and the detailed clinical information available in some studies, wherein binary, severity-encompassing, and "precision," data-driven clinical traits are employed. As an illustration of the use of data-driven traits across multiple cohorts, we present an application of dental caries experience data harmonization in 8 participating studies (N = 55,143) using previously developed permanent dentition tooth surface-level dental caries pattern traits. We demonstrate that these clinical patterns are transferable across multiple cohorts, have similar relative contributions within each study, and thus are prime targets for genetic interrogation in the expanded and diverse multiethnic sample of GLIDE2. We anticipate that results from GLIDE2 will decisively advance the knowledge base of mechanisms at play in oral, dental, and craniofacial health and disease and further catalyze international collaboration and data and resource sharing in genomics research.
Keywords: Genetics; data sciences; dental caries; dentition, permanent; epidemiology; genome-wide association study.
Conflict of interest statement
Figures
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

