Non-volatile signals and redox mechanisms are required for the responses of Arabidopsis roots to Pseudomonas oryzihabitans
- PMID: 36001048
- PMCID: PMC10277831
- DOI: 10.1093/jxb/erac346
Non-volatile signals and redox mechanisms are required for the responses of Arabidopsis roots to Pseudomonas oryzihabitans
Abstract
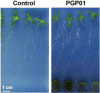
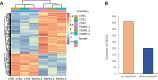
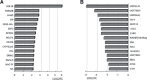
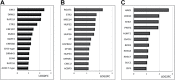
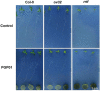
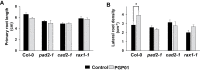
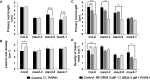
Soil bacteria promote plant growth and protect against environmental stresses, but the mechanisms involved remain poorly characterized, particularly when there is no direct contact between the roots and bacteria. Here, we explored the effects of Pseudomonas oryzihabitans PGP01 on the root system architecture (RSA) in Arabidopsis thaliana seedlings. Significant increases in lateral root (LR) density were observed when seedlings were grown in the presence of P. oryzihabitans, as well as an increased abundance of transcripts associated with altered nutrient transport and phytohormone responses. However, no bacterial transcripts were detected on the root samples by RNAseq analysis, demonstrating that the bacteria do not colonize the roots. Separating the agar containing bacteria from the seedlings prevented the bacteria-induced changes in RSA. Bacteria-induced changes in RSA were absent from mutants defective in ethylene response factor (ERF109), glutathione synthesis (pad2-1, cad2-1, and rax1-1) and in strigolactone synthesis (max3-9 and max4-1) or signalling (max2-3). However, the P. oryzihabitans-induced changes in RSA were similar in the low ascorbate mutants (vtc2-1and vtc2-2) to the wild-type controls. Taken together, these results demonstrate the importance of non-volatile signals and redox mechanisms in the root architecture regulation that occurs following long-distance perception of P. oryzihabitans.
Keywords: Pseudomonas oryzihabitans; Ascorbate; ethylene-responsive transcription factor 109; glutathione; plant growth-promoting rhizobacteria; reactive oxygen species; root system architecture.
© The Author(s) 2022. Published by Oxford University Press on behalf of the Society for Experimental Biology. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Conflict of interest statement
The authors have no conflicts to declare.
Figures

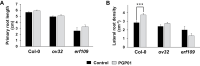


References
-
- Ahemad M, Kibret M. 2014. Mechanisms and applications of plant growth promoting rhizobacteria: current perspective. Journal of King Saud University-Science 26, 1–20.
-
- Akiyama K, Matsuzaki K, Hayashi H. 2005. Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435, 824–827. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

