Fungi-on-a-Chip: microfluidic platforms for single-cell studies on fungi
- PMID: 36001464
- PMCID: PMC9779915
- DOI: 10.1093/femsre/fuac039
Fungi-on-a-Chip: microfluidic platforms for single-cell studies on fungi
Abstract
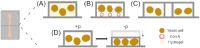
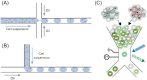
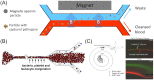
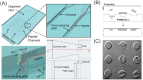
This review highlights new advances in the emerging field of 'Fungi-on-a-Chip' microfluidics for single-cell studies on fungi and discusses several future frontiers, where we envisage microfluidic technology development to be instrumental in aiding our understanding of fungal biology. Fungi, with their enormous diversity, bear essential roles both in nature and our everyday lives. They inhabit a range of ecosystems, such as soil, where they are involved in organic matter degradation and bioremediation processes. More recently, fungi have been recognized as key components of the microbiome in other eukaryotes, such as humans, where they play a fundamental role not only in human pathogenesis, but also likely as commensals. In the food sector, fungi are used either directly or as fermenting agents and are often key players in the biotechnological industry, where they are responsible for the production of both bulk chemicals and antibiotics. Although the macroscopic fruiting bodies are immediately recognizable by most observers, the structure, function, and interactions of fungi with other microbes at the microscopic scale still remain largely hidden. Herein, we shed light on new advances in the emerging field of Fungi-on-a-Chip microfluidic technologies for single-cell studies on fungi. We discuss the development and application of microfluidic tools in the fields of medicine and biotechnology, as well as in-depth biological studies having significance for ecology and general natural processes. Finally, a future perspective is provided, highlighting new frontiers in which microfluidic technology can benefit this field.
Keywords: Fungi-on-a-Chip microfluidic technology; arbuscular mycorrhizal fungi; fungal biology; fungal highways; single-cell microscopy; yeast.
© The Author(s) 2022. Published by Oxford University Press on behalf of FEMS.
Figures








Similar articles
-
Microbiome-on-a-Chip: New Frontiers in Plant-Microbiota Research.Trends Microbiol. 2017 Aug;25(8):610-613. doi: 10.1016/j.tim.2017.05.001. Epub 2017 May 24. Trends Microbiol. 2017. PMID: 28550943
-
Microfluidics as an Emerging Platform for Exploring Soil Environmental Processes: A Critical Review.Environ Sci Technol. 2022 Jan 18;56(2):711-731. doi: 10.1021/acs.est.1c03899. Epub 2022 Jan 5. Environ Sci Technol. 2022. PMID: 34985862 Review.
-
Microfluidic Tools for Probing Fungal-Microbial Interactions at the Cellular Level.J Vis Exp. 2022 Jun 23;(184). doi: 10.3791/63917. J Vis Exp. 2022. PMID: 35815981
-
Microfluidics in Biotechnology: Overview and Status Quo.Adv Biochem Eng Biotechnol. 2022;179:1-16. doi: 10.1007/10_2022_206. Adv Biochem Eng Biotechnol. 2022. PMID: 35333948 Review.
-
Soil-on-a-Chip: microfluidic platforms for environmental organismal studies.Lab Chip. 2016 Jan 21;16(2):228-41. doi: 10.1039/c5lc01285f. Lab Chip. 2016. PMID: 26645910 Review.
Cited by
-
Studying root-environment interactions in structured microdevices.J Exp Bot. 2023 Jul 18;74(13):3851-3863. doi: 10.1093/jxb/erad122. J Exp Bot. 2023. PMID: 37042515 Free PMC article.
-
Reliable replicative lifespan determination of yeast with a single-channel microfluidic chip.Biol Open. 2024 Nov 15;13(11):bio060596. doi: 10.1242/bio.060596. Epub 2024 Nov 26. Biol Open. 2024. PMID: 39479938 Free PMC article.
-
Aging on Chip: Harnessing the Potential of Microfluidic Technologies in Aging and Rejuvenation Research.Adv Healthc Mater. 2025 Aug;14(20):e2500217. doi: 10.1002/adhm.202500217. Epub 2025 Jun 12. Adv Healthc Mater. 2025. PMID: 40509615 Free PMC article. Review.
-
AMF-SporeChip provides new insights into arbuscular mycorrhizal fungal asymbiotic hyphal growth dynamics at the cellular level.Lab Chip. 2024 Mar 26;24(7):1930-1946. doi: 10.1039/d3lc00859b. Lab Chip. 2024. PMID: 38416560 Free PMC article.
-
Monitoring the impact of confinement on hyphal penetration and fungal behavior.PLoS One. 2024 Dec 30;19(12):e0312855. doi: 10.1371/journal.pone.0312855. eCollection 2024. PLoS One. 2024. PMID: 39775212 Free PMC article.
References
-
- Aleklett K, Boddy L.. Fungal behaviour: a new frontier in behavioural ecology. Trends Ecol Evol. 2021;36:787–96. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

