Genomic profiling for clinical decision making in lymphoid neoplasms
- PMID: 36001803
- PMCID: PMC9837456
- DOI: 10.1182/blood.2022015854
Genomic profiling for clinical decision making in lymphoid neoplasms
Abstract
With the introduction of large-scale molecular profiling methods and high-throughput sequencing technologies, the genomic features of most lymphoid neoplasms have been characterized at an unprecedented scale. Although the principles for the classification and diagnosis of these disorders, founded on a multidimensional definition of disease entities, have been consolidated over the past 25 years, novel genomic data have markedly enhanced our understanding of lymphomagenesis and enriched the description of disease entities at the molecular level. Yet, the current diagnosis of lymphoid tumors is largely based on morphological assessment and immunophenotyping, with only few entities being defined by genomic criteria. This paper, which accompanies the International Consensus Classification of mature lymphoid neoplasms, will address how established assays and newly developed technologies for molecular testing already complement clinical diagnoses and provide a novel lens on disease classification. More specifically, their contributions to diagnosis refinement, risk stratification, and therapy prediction will be considered for the main categories of lymphoid neoplasms. The potential of whole-genome sequencing, circulating tumor DNA analyses, single-cell analyses, and epigenetic profiling will be discussed because these will likely become important future tools for implementing precision medicine approaches in clinical decision making for patients with lymphoid malignancies.
© 2022 by The American Society of Hematology.
Conflict of interest statement
Conflict-of-interest disclosure: L.d.L. reports consultancy for AbbVie, Bayer, Bio Ascend, Lunaphore, Novartis. A.A.A. reports consultancy for Celgene, Chugai, Genentech, Gilead, Janssen, Pharmacyclics, and Roche; research funding from Celgene and Pfizer; and ownership interests CiberMed, ForeSight, and FortySeven. E.C. reports consultancy for Genmab, Illumina, NanoString, and Takeda; educational honoraria from AstraZeneca, EUSA Pharma, Janssen, and Takeda; is a named inventor on a patent describing the use of gene expression to subtype aggressive B-cell lymphomas, one of which is licensed to NanoString Technologies, and an author in a protected bioinformatic pipeline (IgCaller) licensed to Diagnostica Longwood. A. Dogan reports consultancy for EUSA Pharma, Incyte, and Loxo; and research funding from Roche and Takeda. S.M.H. reports consultancy for Cimieo Therapeutics, Daiichi Sankyo, Kyowa Hakko Kirin, ONO Pharmaceuticals, SecuraBio, Shoreline Biosciences, Takeda, Tubulis, and Yingli Pharma; and research funding from ADC Therapeutics, Affimed, C4, Celgene, Crispr Therapeutics, Daiichi Sankyo, Kyowa Hakko Kirin, Millennium/Takeda, Seattle Genetics, and Verastem/SecuraBio. A.M.M. reports consulting for AstraZeneca, Epizyme, Exo Therapeutics, Janssen, and Treeline Biosciences; and research funding from AstraZeneca, Epizyme, and Janssen. R.R. reports consultancy for AbbVie, AstraZeneca, Illumina, Janssen, and Roche. D.R. reports consultancy for AbbVie, AstraZeneca, BeiGene, BMS, and Janssen; and research funding from AbbVie, AstraZeneca, BeiGene, BMS, and Janssen. C.S. reports consultancy for AbbVie, Bayer, and Seattle Genetics; and research funding from Trillium Therapeutics, BMS, and Epizyme. S.P.T. reports consultancy for BeiGene, Janssen, and Pharmacyclics; and research funding from BeiGene, BMS, Eli Lilly, Janssen, Pharmacyclics, and X4 Pharmaceuticals. A.D.Z. reports consultancy for Amgen, AstraZeneca, BeiGene, BMS/Celgene/JUNO, Genentech/Roche, Gilead/Kite, Janssen, MEI Pharma, Novartis, and Pharmacyclics/AbbVie; scientific advisory board for Adaptive Biotechnologies and Lymphoma Research Foundation; and research funding from AbbVie, BeiGene, Genentech/Roche, and MEI Pharma. C.E.A. reports consultancy for Electra and Sobi; and research support from Roche/Genetech. W.C.C. is a named inventor on a patent describing the use of gene expression to subtype aggressive B-cell lymphomas, one of which is licensed to NanoString Technologies. L.C. reports advisory boards for AbbVie and Roche. F.d’A. reports advisory board for Kyova Kirin; and research funding from Servier. S.D. reports consultancy for Incyte, Roche, and Takeda. M.D. reports speakers’ honoraria from Amgen, AstraZeneca, Bayer, BMS/Celgene, Gilead/Kite, Incyte, Janssen, Novartis, and Roche; scientific advisory board for AstraZeneca, Bayer, BMS/Celgene, Genmab, Gilead/Kite, Incyte, Janssen, Lilly/Loxo, MorphoSys, Novartis, and Roche; and research funding (institutional) from AbbVie, Bayer, BMS/Celgene, Gilead/Kite, Janssen, and Roche. K.D. reports scientific advisory board for AbbVie, ADC Therapeutics, AstraZeneca, BeiGene, Daiichi Sankyo, Genentech, Genmab, Incyte, and MorphoSys. A.L.F. reports research funding from Seattle Genetics; and is a named inventor on technology for which Mayo Clinic holds unlicensed patents. F.F. reports advisory boards or speaker honoraria from EUROPharma, Roche, and Stemline; and research funding from Stemline. P. Gaulard reports consultancy for Gilead and Takeda; and research funding from Alderan, Innate Pharma, Sanofi, and Takeda. P. Ghia reports consultancy for AbbVie, AstraZeneca, BeiGene, BMS, Janssen, Lilly/Loxo, MSD, and Roche; and research funding from AbbVie, AstraZeneca, and Janssen. J.G.G. reports honoraria from AbbVie, Amgen, AstraZeneca, BMS, Gilead/Kite, Janssen, and Novartis; and research funding from AstraZeneca, BMS, and Janssen. O.H. reports holding stock in AB science; is a cofounder of AB science and Inatherys; and reports research funding from AB science, AbbVie, Alexion, BMS, Inatherys, Innate Pharma, and Takeda. D.J.H. reports research funding from AstraZeneca. E.D.H. reports consultancy for Abcon, Astellas, Cytomx, and Novartis; and research funding from AbbVie, Eli Lilly, and Virtuoso. K. Karube reports speaker honoraria from AstraZeneca, Chugai Pharmaceutical, Eisai, Kyowa Kirin, Janssen, Otsuka Pharmaceutical, Takeda, Symbio, and Meiji Seika Pharma; and research funding from Takeda. K. Kataoka reports honoraria from Astellas Pharma, AstraZeneca, Chugai Pharmaceutical, Eisai, Kyowa Kirin, Janssen, Novartis, Ono Pharmaceutical, Otsuka Pharmaceutical, Sumitomo Dainippon Pharma, and Takeda; and research funding from Asahi Kasei Pharma, Chugai Pharmaceutical, Chordia Therapeutics, Eisai, Kyowa Kirin, Japan Blood Products Organization, JCR Pharmaceutical, Mochida Pharmaceutical, Nippon Shinyaku, Ono Pharmaceutical, Otsuka Pharmaceutical, Shionogi, Sumitomo Dainippon Pharmaceutical, Takeda, and Teijin Pharma; is a named inventor on a patent for genetic alterations as a biomarker in T-cell lymphomas and for PD-L1 abnormalities as a predictive biomarker for immune checkpoint blockade therapy; and reports stocks in Asahi Genomics. W.S.K. reports research funding from BeiGene, Boryong, Kyowa Kirin, Merck, Roche, and Sanofi. G.L. reports consultancy for AbbVie, ADC, AstraZeneca, Bayer, BMS, Celgene, Constellation, Gilead, Genmab, Incyte, Janssen, Karyopharm, MorphoSys, NanoString, Novartis, Roche, and Takeda; speakers’ honoraria from AbbVie, Bayer, Celgene, Janssen, and Roche; and research funding from AstraZeneca, Bayer, Celgene, Gilead, Janssen, MorphoSys, Novartis, and Roche. L.P. reports research funding from AstraZeneca. M.A.P. reports speaker honoraria and advisory board fees from Celgene, Gilead, Janssen, Kyowa Kirin, Millennium/Takeda, and NanoString. S.J.R. reports research funding from BMS and Merck. G.A.S. reports consulting and advisory boards for AbbVie, Bayer, BeiGene, BMS/Celgene, Epizyme, Genentech/Roche, Genmab, Incyte, Janssen, Kite/Gilead, Loxo, Miltenyi, Molecular Partners, MorphoSys, Nordic Nanovector, Novartis, Rapt, Regeneron, and Takeda; and is a shareholder of Owkin. J.S.-M. reports consultancy and advisory boards for AbbVie, Amgen, BMS, Celgene, GSK, Haemalogix, Janssen-Cilag, Karyopharm, MSD, Novartis, Regeneron, Roche, Sanofi, SecuraBio, and Takeda. K.J.S. reports consultancy for BMS, Janssen, Kyowa, Merck, Novartis, and Seattle Genetics; and research funding from BMS. L.H.S. reports consulting and honoraria from AbbVie, Acerta, Amgen, Apbiologix, AstraZeneca, Celgene, Chugai, Gilead, Incyte, Janssen, Kite, Karyopharm, Lundbeck, Merck, MorphoSys, Roche/Genentech, Sandoz, Seattle Genetics, Servier, Takeda, Teva, TG Therapeutics, and Verastem; and research funding from Roche/Genentech and Teva. L.M.S. is one of the developers of the LymphGen software, and NCI has applied for copyright protection for this software; and is an inventor on NCI patents relevant to cell-of-origin classification of DLBCL. C.S.T. reports honoraria from AbbVie, BeiGene, and Janssen. J.T. reports research funds (institutional) from BeiGene, BMS, Cellectar, Janssen, Pharmacyclics, Roche, and Takeda. C.J.W. reports equity in BioNTech; research funds from Pharmacyclics. P.L.Z. reports consultancy for MSD TG Therapeutics, EUSAPharma, and Novartis; advisory board and speakers’ bureau for BeiGene, BMS, Celltrion, EUSAPharma, Gilead, Incyte, Janssen-Cilag, Kyowa Kirin, MSD TG Therapeutics, Novartis, Roche, Servier, and Takeda; and advisory board for ADC Therapeutics, Sandoz, and Secura Bio. D.W.S. reports consultancy for AbbVie, AstraZeneca, Incyte, and Janssen; research funds from Janssen and Roche/Genentech; and is a named inventor on a patent describing the use of gene expression to subtype aggressive B-cell lymphomas, one of which is licensed to NanoString Technologies. The remaining authors declare no competing financial interests.
Figures

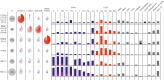

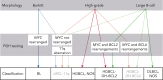


References
-
- Swerdlow SH, Campo E, Harris NL, et al., editors. WHO Classification of Tumours of the Haematopoietic and Lymphoid Tissues. Revised 4th ed. IARC; 2017.
-
- Zech L, Haglund U, Nilsson K, Klein G. Characteristic chromosomal abnormalities in biopsies and lymphoid-cell lines from patients with Burkitt and non-Burkitt lymphomas. Int J Cancer. 1976;17:47–56. - PubMed
-
- van den Brand M, Rijntjes J, Mobs M, et al. Next-generation sequencing-based clonality assessment of Ig gene rearrangements: a multicenter validation study by EuroClonality-NGS. J Mol Diagn. 2021;23(9):1105–1115. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

