Advances in Computational Polypharmacology
- PMID: 36002382
- PMCID: PMC10078381
- DOI: 10.1002/minf.202200190
Advances in Computational Polypharmacology
Abstract
In drug discovery, polypharmacology encompasses the use of small molecules with defined multi-target activity and in vivo effects resulting from multi-target engagement. Multi-target compounds are often efficacious in the treatment of complex diseases involving target and pathway networks, but might also elicit unwanted side effects. Computational approaches such as target prediction or multi-target ligand design have been used to support polypharmacological drug discovery. In addition to efforts directed at the identification or design of new multi-target compounds, other computational investigations have aimed to differentiate such compounds from potential false-positives or explore the molecular basis of multi-target activities. Herein, a concise overview of the field is provided and recent advances in computational polypharmacology through machine learning are discussed.
Keywords: Polypharmacology; computational methods; explainable machine learning; medicinal chemistry; molecular design; multi-target compounds.
© 2022 The Authors. Molecular Informatics published by Wiley-VCH GmbH.
Conflict of interest statement
None declared.
Figures
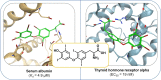


References
-
- J. U. Peters (Ed.), Polypharmacology in Drug Discovery. John Wiley & Sons, Hoboken, 2012.
-
- Anighoro A., Bajorath J., Rastelli G., J. Med. Chem. 2014, 57, 7874–7887. - PubMed
-
- Benek O., Korabecny J., Soukup O., Trends Pharmacol. Sci. 2020, 41, 434–445. - PubMed
-
- Tschöp M. H., Finan B., Clemmensen C., Gelfanov V., Perez-Tilve D., Müller T. D., DiMarchi R. D., Cell Metab. 2016, 24, 51–62. - PubMed
Publication types
LinkOut - more resources
Full Text Sources

