R-loop formation and conformational activation mechanisms of Cas9
- PMID: 36002571
- PMCID: PMC9433323
- DOI: 10.1038/s41586-022-05114-0
R-loop formation and conformational activation mechanisms of Cas9
Erratum in
-
Publisher Correction: R-loop formation and conformational activation mechanisms of Cas9.Nature. 2023 Nov;623(7987):E10. doi: 10.1038/s41586-023-06779-x. Nature. 2023. PMID: 37903882 Free PMC article. No abstract available.
Abstract
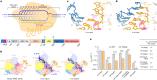
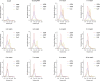
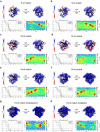
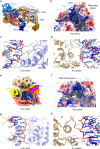
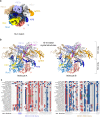
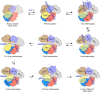
Cas9 is a CRISPR-associated endonuclease capable of RNA-guided, site-specific DNA cleavage1-3. The programmable activity of Cas9 has been widely utilized for genome editing applications4-6, yet its precise mechanisms of target DNA binding and off-target discrimination remain incompletely understood. Here we report a series of cryo-electron microscopy structures of Streptococcus pyogenes Cas9 capturing the directional process of target DNA hybridization. In the early phase of R-loop formation, the Cas9 REC2 and REC3 domains form a positively charged cleft that accommodates the distal end of the target DNA duplex. Guide-target hybridization past the seed region induces rearrangements of the REC2 and REC3 domains and relocation of the HNH nuclease domain to assume a catalytically incompetent checkpoint conformation. Completion of the guide-target heteroduplex triggers conformational activation of the HNH nuclease domain, enabled by distortion of the guide-target heteroduplex, and complementary REC2 and REC3 domain rearrangements. Together, these results establish a structural framework for target DNA-dependent activation of Cas9 that sheds light on its conformational checkpoint mechanism and may facilitate the development of novel Cas9 variants and guide RNA designs with enhanced specificity and activity.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







Similar articles
-
Cryo-EM structures reveal coordinated domain motions that govern DNA cleavage by Cas9.Nat Struct Mol Biol. 2019 Aug;26(8):679-685. doi: 10.1038/s41594-019-0258-2. Epub 2019 Jul 8. Nat Struct Mol Biol. 2019. PMID: 31285607 Free PMC article.
-
Coordinated Actions of Cas9 HNH and RuvC Nuclease Domains Are Regulated by the Bridge Helix and the Target DNA Sequence.Biochemistry. 2021 Dec 14;60(49):3783-3800. doi: 10.1021/acs.biochem.1c00354. Epub 2021 Nov 10. Biochemistry. 2021. PMID: 34757726 Free PMC article.
-
The initiation, propagation and dynamics of CRISPR-SpyCas9 R-loop complex.Nucleic Acids Res. 2018 Jan 9;46(1):350-361. doi: 10.1093/nar/gkx1117. Nucleic Acids Res. 2018. PMID: 29145633 Free PMC article.
-
Allosteric regulation of CRISPR-Cas9 for DNA-targeting and cleavage.Curr Opin Struct Biol. 2020 Jun;62:166-174. doi: 10.1016/j.sbi.2020.01.013. Epub 2020 Feb 18. Curr Opin Struct Biol. 2020. PMID: 32070859 Free PMC article. Review.
-
CRISPR-Cas9 Structures and Mechanisms.Annu Rev Biophys. 2017 May 22;46:505-529. doi: 10.1146/annurev-biophys-062215-010822. Epub 2017 Mar 30. Annu Rev Biophys. 2017. PMID: 28375731 Review.
Cited by
-
Gene editing of angiotensin for blood pressure management.Int J Cardiol Cardiovasc Risk Prev. 2024 Aug 20;23:200323. doi: 10.1016/j.ijcrp.2024.200323. eCollection 2024 Dec. Int J Cardiol Cardiovasc Risk Prev. 2024. PMID: 39258007 Free PMC article.
-
Insights into the Mechanism of CRISPR/Cas9-Based Genome Editing from Molecular Dynamics Simulations.ACS Omega. 2022 Dec 30;8(2):1817-1837. doi: 10.1021/acsomega.2c05583. eCollection 2023 Jan 17. ACS Omega. 2022. PMID: 36687047 Free PMC article. Review.
-
Quantifying Protein-Nucleic Acid Interactions for Engineering Useful CRISPR-Cas9 Genome-Editing Variants.Methods Mol Biol. 2025;2870:227-243. doi: 10.1007/978-1-0716-4213-9_12. Methods Mol Biol. 2025. PMID: 39543038
-
CRISPR-Cas12a bends DNA to destabilize base pairs during target interrogation.bioRxiv [Preprint]. 2024 Jul 31:2024.07.31.606079. doi: 10.1101/2024.07.31.606079. bioRxiv. 2024. Update in: Nucleic Acids Res. 2025 Jan 11;53(2):gkae1192. doi: 10.1093/nar/gkae1192. PMID: 39131396 Free PMC article. Updated. Preprint.
-
Cleavage of DNA Substrate Containing Nucleotide Mismatch in the Complementary Region to sgRNA by Cas9 Endonuclease: Thermodynamic and Structural Features.Int J Mol Sci. 2024 Oct 9;25(19):10862. doi: 10.3390/ijms251910862. Int J Mol Sci. 2024. PMID: 39409191 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

