Modeling of pneumococcal serogroup 10 capsular polysaccharide molecular conformations provides insight into epitopes and observed cross-reactivity
- PMID: 36003080
- PMCID: PMC9393222
- DOI: 10.3389/fmolb.2022.961532
Modeling of pneumococcal serogroup 10 capsular polysaccharide molecular conformations provides insight into epitopes and observed cross-reactivity
Abstract
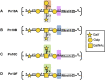
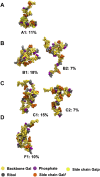
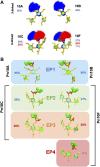
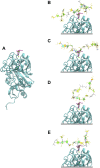
Streptococcus pneumoniae is an encapsulated gram-negative bacterium and a significant human pathogen. The capsular polysaccharide (CPS) is essential for virulence and a target antigen for vaccines. Although widespread introduction of pneumococcal conjugate vaccines (PCVs) has significantly reduced disease, the prevalence of non-vaccine serotypes has increased. On the basis of the CPS, S. pneumoniae serogroup 10 comprises four main serotypes 10A, 10B, 10C, and 10F; as well as the recently identified 10D. As it is the most prevalent, serotype 10A CPS has been included as a vaccine antigen in the next generation PCVs. Here we use molecular modeling to provide conformational rationales for the complex cross-reactivity reported between serotypes 10A, 10B, 10C, and 10F anti-sera. Although the highly mobile phosphodiester linkages produce very flexible CPS, shorter segments are conformationally defined, with exposed -D-galactofuranose ( DGalf) side chains that are potential antibody binding sites. We identify four distinct conformational epitopes for the immunodominant DGalf that assist in rationalizing the complex asymmetric cross-reactivity relationships. In particular, we find that strongly cross-reactive serotypes share common epitopes. Further, we show that human intelectin-1 has the potential to bind the exposed exocyclic 1,2-diol of the terminal DGalf in each serotype; the relative accessibility of three- or six-linked DGalf may play a role in the strength of the innate immune response and hence serotype disease prevalence. In conclusion, our modeling study and relevant serological studies support the inclusion of serotype 10A in a vaccine to best protect against serogroup 10 disease.
Keywords: S. pneumoniae; capsular polysaccharide; carbohydrate epitopes; cross-protection; molecular modeling; serogroup 10; vaccine antigen.
Copyright © 2022 Richardson, Kuttel and Ravenscroft.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
A New Pneumococcal Capsule Type, 10D, is the 100th Serotype and Has a Large cps Fragment from an Oral Streptococcus.mBio. 2020 May 19;11(3):e00937-20. doi: 10.1128/mBio.00937-20. mBio. 2020. PMID: 32430472 Free PMC article.
-
Nonpneumococcal Strains Recently Recovered from Carriage Specimens and Expressing Capsular Serotypes Highly Related or Identical to Pneumococcal Serotypes 2, 4, 9A, 13, and 23A.mBio. 2021 May 18;12(3):e01037-21. doi: 10.1128/mBio.01037-21. mBio. 2021. PMID: 34006665 Free PMC article.
-
Comparative Molecular Modelling of Capsular Polysaccharide Conformations in Streptococcus suis Serotypes 1, 2, 1/2 and 14 Identifies Common Epitopes for Antibody Binding.Front Mol Biosci. 2022 Feb 8;9:830854. doi: 10.3389/fmolb.2022.830854. eCollection 2022. Front Mol Biosci. 2022. PMID: 35211512 Free PMC article.
-
A systematic review of invasive pneumococcal disease vaccine failures and breakthrough with higher-valency pneumococcal conjugate vaccines in children.Expert Rev Vaccines. 2022 Feb;21(2):201-214. doi: 10.1080/14760584.2022.2012455. Epub 2022 Feb 3. Expert Rev Vaccines. 2022. PMID: 34882050
-
Relationship between immune response to pneumococcal conjugate vaccines in infants and indirect protection after vaccine implementation.Expert Rev Vaccines. 2019 Jun;18(6):641-661. doi: 10.1080/14760584.2019.1627207. Epub 2019 Jun 22. Expert Rev Vaccines. 2019. PMID: 31230486 Review.
Cited by
-
Streptococcus pneumoniae serotype 33G: genetic, serological, and structural analysis of a new capsule type.Microbiol Spectr. 2024 Jan 11;12(1):e0357923. doi: 10.1128/spectrum.03579-23. Epub 2023 Dec 7. Microbiol Spectr. 2024. PMID: 38059623 Free PMC article.
-
Strengths and weaknesses of pneumococcal conjugate vaccines.Glycoconj J. 2023 Apr;40(2):135-148. doi: 10.1007/s10719-023-10100-3. Epub 2023 Jan 18. Glycoconj J. 2023. PMID: 36652051 Free PMC article. Review.
-
Conformational comparisons of Pasteurella multocida types B and E and structurally related capsular polysaccharides.Glycobiology. 2023 Oct 29;33(9):745-754. doi: 10.1093/glycob/cwad049. Glycobiology. 2023. PMID: 37334939 Free PMC article.
References
-
- Bush C. A., Yang J., Yu B., Cisar J. O. (2014). Chemical structures of Streptococcus pneumoniae capsular polysaccharide type 39 (CPS39), CPS47F, and CPS34 characterized by nuclear magnetic resonance spectroscopy and their relation to CPS10A. J. Bacteriol. 196 (18), 3271–3278. 10.1128/JB.01731-14 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

