Drosophila melanogaster: A platform for anticancer drug discovery and personalized therapies
- PMID: 36003330
- PMCID: PMC9393232
- DOI: 10.3389/fgene.2022.949241
Drosophila melanogaster: A platform for anticancer drug discovery and personalized therapies
Abstract
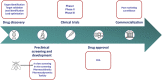
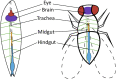
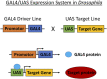
Cancer is a complex disease whereby multiple genetic aberrations, epigenetic modifications, metabolic reprogramming, and the microenvironment contribute to the development of a tumor. In the traditional anticancer drug discovery pipeline, drug candidates are usually screened in vitro using two-dimensional or three-dimensional cell culture. However, these methods fail to accurately mimic the human disease state. This has led to the poor success rate of anticancer drugs in the preclinical stages since many drugs are abandoned due to inefficacy or toxicity when transitioned to whole-organism models. The common fruit fly, Drosophila melanogaster, has emerged as a beneficial system for modeling human cancers. Decades of fundamental research have shown the evolutionary conservation of key genes and signaling pathways between flies and humans. Moreover, Drosophila has a lower genetic redundancy in comparison to mammals. These factors, in addition to the advancement of genetic toolkits for manipulating gene expression, allow for the generation of complex Drosophila genotypes and phenotypes. Numerous studies have successfully created Drosophila models for colorectal, lung, thyroid, and brain cancers. These models were utilized in the high-throughput screening of FDA-approved drugs which led to the identification of several compounds capable of reducing proliferation and rescuing phenotypes. More noteworthy, Drosophila has also unlocked the potential for personalized therapies. Drosophila 'avatars' presenting the same mutations as a patient are used to screen multiple therapeutic agents targeting multiple pathways to find the most appropriate combination of drugs. The outcomes of these studies have translated to significant responses in patients with adenoid cystic carcinoma and metastatic colorectal cancers. Despite not being widely utilized, the concept of in vivo screening of drugs in Drosophila is making significant contributions to the current drug discovery pipeline. In this review, we discuss the application of Drosophila as a platform in anticancer drug discovery; with special focus on the cancer models that have been generated, drug libraries that have been screened and the status of personalized therapies. In addition, we elaborate on the biological and technical limitations of this system.
Keywords: Drosophila melanogaster; cancer models; drug discovery; high-throughput screening; personalized therapy.
Copyright © 2022 Munnik, Xaba, Malindisa, Russell and Sooklal.
Conflict of interest statement
BR is funded by the company Buboo (Pty) Ltd. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
References
-
- Aird R. E., Cummings J., Ritchie A. A., Muir M., Morris R. E., Chen H., et al. (2002). In vitro and in vivo activity and cross resistance profiles of novel ruthenium (II) organometallic arene complexes in human ovarian cancer. Br. J. Cancer 86 (10), 1652–1657. 10.1038/sj.bjc.6600290 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

