Pervasive Listeria monocytogenes Is Common in the Norwegian Food System and Is Associated with Increased Prevalence of Stress Survival and Resistance Determinants
- PMID: 36005805
- PMCID: PMC9499026
- DOI: 10.1128/aem.00861-22
Pervasive Listeria monocytogenes Is Common in the Norwegian Food System and Is Associated with Increased Prevalence of Stress Survival and Resistance Determinants
Abstract
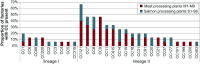
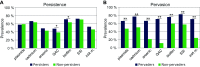
To investigate the diversity, distribution, persistence, and prevalence of stress survival and resistance genes of Listeria monocytogenes clones dominating in food processing environments in Norway, genome sequences from 769 L. monocytogenes isolates from food industry environments, foods, and raw materials (512 of which were sequenced in the present study) were subjected to whole-genome multilocus sequence typing (wgMLST), single-nucleotide polymorphism (SNP), and comparative genomic analyses. The data set comprised isolates from nine meat and six salmon processing facilities in Norway collected over a period of three decades. The most prevalent clonal complex (CC) was CC121, found in 10 factories, followed by CC7, CC8, and CC9, found in 7 factories each. Overall, 72% of the isolates were classified as persistent, showing 20 or fewer wgMLST allelic differences toward an isolate found in the same factory in a different calendar year. Moreover, over half of the isolates (56%) showed this level of genetic similarity toward an isolate collected from a different food processing facility. These were designated as pervasive strains, defined as clusters with the same level of genetic similarity as persistent strains but isolated from different factories. The prevalence of genetic determinants associated with increased survival in food processing environments, including heavy metal and biocide resistance determinants, stress response genes, and inlA truncation mutations, showed a highly significant increase among pervasive isolates but not among persistent isolates. Furthermore, these genes were significantly more prevalent among the isolates from food processing environments compared to in isolates from natural and rural environments (n = 218) and clinical isolates (n = 111) from Norway. IMPORTANCE Listeria monocytogenes can persist in food processing environments for months to decades and spread through the food system by, e.g., contaminated raw materials. Knowledge of the distribution and diversity of L. monocytogenes is important in outbreak investigations and is essential to effectively track and control this pathogen in the food system. The present study presents a comprehensive overview of the prevalence of persistent clones and of the diversity of L. monocytogenes in Norwegian food processing facilities. The results demonstrate extensive spread of highly similar strains throughout the Norwegian food system, in that 56% of the 769 collected isolates from food processing factories belonged to clusters of L. monocytogenes identified in more than one facility. These strains were associated with an overall increase in the prevalence of plasmids and determinants of heavy metal and biocide resistance, as well as other genetic elements associated with stress survival mechanisms and persistence.
Keywords: Listeria monocytogenes; WGS; antibiotic resistance; food processing environment; food safety; inlA; meat industry; meat processing; persistence; pervasive; plasmids; salmon industry; salmon processing; stress resistance; stress survival; whole-genome sequencing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Whole-Genome Sequencing Analysis of Listeria monocytogenes from Rural, Urban, and Farm Environments in Norway: Genetic Diversity, Persistence, and Relation to Clinical and Food Isolates.Appl Environ Microbiol. 2022 Mar 22;88(6):e0213621. doi: 10.1128/aem.02136-21. Epub 2022 Feb 2. Appl Environ Microbiol. 2022. PMID: 35108102 Free PMC article.
-
In-Depth Longitudinal Study of Listeria monocytogenes ST9 Isolates from the Meat Processing Industry: Resolving Diversity and Transmission Patterns Using Whole-Genome Sequencing.Appl Environ Microbiol. 2020 Jul 2;86(14):e00579-20. doi: 10.1128/AEM.00579-20. Print 2020 Jul 2. Appl Environ Microbiol. 2020. PMID: 32414794 Free PMC article.
-
Unraveling the emergence and population diversity of Listeria monocytogenes in a newly built meat facility through whole genome sequencing.Int J Food Microbiol. 2021 Feb 16;340:109043. doi: 10.1016/j.ijfoodmicro.2021.109043. Epub 2021 Jan 4. Int J Food Microbiol. 2021. PMID: 33454520
-
Biocide use in the food industry and the disinfectant resistance of persistent strains of Listeria monocytogenes and Escherichia coli.Symp Ser Soc Appl Microbiol. 2002;(31):111S-120S. Symp Ser Soc Appl Microbiol. 2002. PMID: 12481836 Review.
-
Potential Impact of the Resistance to Quaternary Ammonium Disinfectants on the Persistence of Listeria monocytogenes in Food Processing Environments.Front Microbiol. 2016 May 2;7:638. doi: 10.3389/fmicb.2016.00638. eCollection 2016. Front Microbiol. 2016. PMID: 27199964 Free PMC article. Review.
Cited by
-
Reduction and Growth Inhibition of Listeria monocytogenes by Use of Anti-Listerial Nisin, P100 Phages and Buffered Dry Vinegar Fermentates in Standard and Sodium-Reduced Cold-Smoked Salmon.Foods. 2023 Dec 6;12(24):4391. doi: 10.3390/foods12244391. Foods. 2023. PMID: 38137194 Free PMC article.
-
DegU-mediated suppression of carbohydrate uptake in Listeria monocytogenes increases adaptation to oxidative stress.Appl Environ Microbiol. 2023 Oct 31;89(10):e0101723. doi: 10.1128/aem.01017-23. Epub 2023 Oct 3. Appl Environ Microbiol. 2023. PMID: 37787570 Free PMC article.
-
Genetic population structure of Listeria monocytogenes strains isolated from salmon and trout sectors in France.Heliyon. 2023 Jul 10;9(7):e18154. doi: 10.1016/j.heliyon.2023.e18154. eCollection 2023 Jul. Heliyon. 2023. PMID: 37483814 Free PMC article.
-
Genomic characterization of Listeria monocytogenes recovered from dairy facilities in British Columbia, Canada from 2007 to 2017.Front Microbiol. 2024 Mar 22;15:1304734. doi: 10.3389/fmicb.2024.1304734. eCollection 2024. Front Microbiol. 2024. PMID: 38585707 Free PMC article.
-
Whole-Genome Sequence Comparisons of Listeria monocytogenes Isolated from Meat and Fish Reveal High Inter- and Intra-Sample Diversity.Microorganisms. 2022 Oct 26;10(11):2120. doi: 10.3390/microorganisms10112120. Microorganisms. 2022. PMID: 36363712 Free PMC article.
References
-
- Fagerlund A, Idland L, Heir E, Møretrø T, Aspholm M, Lindbäck T, Langsrud S. 2022. WGS analysis of Listeria monocytogenes from rural, urban, and farm environments in Norway: genetic diversity, persistence, and relation to clinical and food isolates. Appl Environ Microbiol 88:e02136-21. 10.1128/aem.02136-21. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

