The small nucleolar RNA SnoR28 regulates plant growth and development by directing rRNA maturation
- PMID: 36005862
- PMCID: PMC9614442
- DOI: 10.1093/plcell/koac265
The small nucleolar RNA SnoR28 regulates plant growth and development by directing rRNA maturation
Abstract
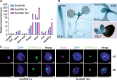
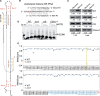
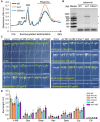
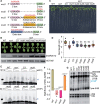
Small nucleolar RNAs (snoRNAs) are noncoding RNAs (ncRNAs) that guide chemical modifications of structural RNAs, which are essential for ribosome assembly and function in eukaryotes. Although numerous snoRNAs have been identified in plants by high-throughput sequencing, the biological functions of most of these snoRNAs remain unclear. Here, we identified box C/D SnoR28.1s as important regulators of plant growth and development by screening a CRISPR/Cas9-generated ncRNA deletion mutant library in Arabidopsis thaliana. Deletion of the SnoR28.1 locus, which contains a cluster of three genes producing SnoR28.1s, resulted in defects in root and shoot growth. SnoR28.1s guide 2'-O-ribose methylation of 25S rRNA at G2396. SnoR28.1s facilitate proper and efficient pre-rRNA processing, as the SnoR28.1 deletion mutants also showed impaired ribosome assembly and function, which may account for the growth defects. SnoR28 contains a 7-bp antisense box, which is required for 2'-O-ribose methylation of 25S rRNA at G2396, and an 8-bp extra box that is complementary to a nearby rRNA methylation site and is partially responsible for methylation of G2396. Both of these motifs are required for proper and efficient pre-rRNA processing. Finally, we show that SnoR28.1s genetically interact with HIDDEN TREASURE2 and NUCLEOLIN1. Our results advance our understanding of the roles of snoRNAs in Arabidopsis.
© American Society of Plant Biologists 2022. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures



Similar articles
-
Identification of 66 box C/D snoRNAs in Arabidopsis thaliana: extensive gene duplications generated multiple isoforms predicting new ribosomal RNA 2'-O-methylation sites.J Mol Biol. 2001 Aug 3;311(1):57-73. doi: 10.1006/jmbi.2001.4851. J Mol Biol. 2001. PMID: 11469857
-
Arabidopsis small nucleolar RNA monitors the efficient pre-rRNA processing during ribosome biogenesis.Proc Natl Acad Sci U S A. 2016 Oct 18;113(42):11967-11972. doi: 10.1073/pnas.1614852113. Epub 2016 Oct 5. Proc Natl Acad Sci U S A. 2016. PMID: 27708161 Free PMC article.
-
Characterisation of the U83 and U84 small nucleolar RNAs: two novel 2'-O-ribose methylation guide RNAs that lack complementarities to ribosomal RNAs.Nucleic Acids Res. 2000 Mar 15;28(6):1348-54. doi: 10.1093/nar/28.6.1348. Nucleic Acids Res. 2000. PMID: 10684929 Free PMC article.
-
Small RNAs with big implications: new insights into H/ACA snoRNA function and their role in human disease.Wiley Interdiscip Rev RNA. 2015 Mar-Apr;6(2):173-89. doi: 10.1002/wrna.1266. Epub 2014 Oct 31. Wiley Interdiscip Rev RNA. 2015. PMID: 25363811 Free PMC article. Review.
-
Small nucleolar RNAs: versatile trans-acting molecules of ancient evolutionary origin.Gene Expr. 2002;10(1-2):17-39. Gene Expr. 2002. PMID: 11868985 Free PMC article. Review.
Cited by
-
Complicated target recognition by archaeal box C/D guide RNAs.Sci China Life Sci. 2024 Apr;67(4):631-644. doi: 10.1007/s11427-022-2412-3. Epub 2023 Nov 30. Sci China Life Sci. 2024. PMID: 38041781
-
Plant ribosomes as a score to fathom the melody of 2'-O-methylation across evolution.RNA Biol. 2024 Jan;21(1):70-81. doi: 10.1080/15476286.2024.2417152. Epub 2024 Nov 7. RNA Biol. 2024. PMID: 39508203 Free PMC article. Review.
-
Unlocking the life code: a review of SnoRNA functional diversity and disease relevance.Cell Commun Signal. 2025 Jun 4;23(1):266. doi: 10.1186/s12964-025-02274-0. Cell Commun Signal. 2025. PMID: 40468441 Free PMC article. Review.
References
-
- Abbasi N, Kim HB, Park NI, Kim HS, Kim YK, Park YI, Choi SB (2010) APUM23, a nucleolar Puf domain protein, is involved in pre-ribosomal RNA processing and normal growth patterning in Arabidopsis. Plant J 64: 960–976 - PubMed
-
- Abel Y, Rederstorff M (2019) SnoRNAs and the emerging class of sdRNAs: multifaceted players in oncogenesis. Biochimie 164: 17–21 - PubMed
-
- Au PC, Helliwell C, Wang MB (2014) Characterizing RNA-protein interaction using cross-linking and metabolite supplemented nuclear RNA-immunoprecipitation. Mol Biol Rep 41: 2971–2977 - PubMed
-
- Bachellerie JP, Cavaille J, Huttenhofer A (2002) The expanding snoRNA world. Biochimie 84: 775–790 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

