The histone deacetylases Rpd3 and Hst1 antagonistically regulate de novo NAD+ metabolism in the budding yeast Saccharomyces cerevisiae
- PMID: 36007612
- PMCID: PMC9486569
- DOI: 10.1016/j.jbc.2022.102410
The histone deacetylases Rpd3 and Hst1 antagonistically regulate de novo NAD+ metabolism in the budding yeast Saccharomyces cerevisiae
Abstract
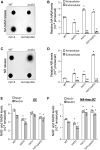
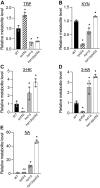
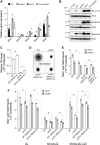
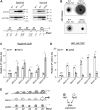
NAD+ is a cellular redox cofactor involved in many essential processes. The regulation of NAD+ metabolism and the signaling networks reciprocally interacting with NAD+-producing metabolic pathways are not yet fully understood. The NAD+-dependent histone deacetylase (HDAC) Hst1 has been shown to inhibit de novo NAD+ synthesis by repressing biosynthesis of nicotinic acid (BNA) gene expression. Here, we alternatively identify HDAC Rpd3 as a positive regulator of de novo NAD+ metabolism in the budding yeast Saccharomyces cerevisiae. We reveal that deletion of RPD3 causes marked decreases in the production of de novo pathway metabolites, in direct contrast to deletion of HST1. We determined the BNA expression profiles of rpd3Δ and hst1Δ cells to be similarly opposed, suggesting the two HDACs may regulate the BNA genes in an antagonistic fashion. Our chromatin immunoprecipitation analysis revealed that Rpd3 and Hst1 mutually influence each other's binding distribution at the BNA2 promoter. We demonstrate Hst1 to be the main deacetylase active at the BNA2 promoter, with hst1Δ cells displaying increased acetylation of the N-terminal tail lysine residues of histone H4, H4K5, and H4K12. Conversely, we show that deletion of RPD3 reduces the acetylation of these residues in an Hst1-dependent manner. This suggests that Rpd3 may function to oppose spreading of Hst1-dependent heterochromatin and represents a unique form of antagonism between HDACs in regulating gene expression. Moreover, we found that Rpd3 and Hst1 also coregulate additional targets involved in other branches of NAD+ metabolism. These findings help elucidate the complex interconnections involved in effecting the regulation of NAD+ metabolism.
Keywords: NAD(+) biosynthesis; cell metabolism; gene regulation; histone deacetylase; metabolic regulation; yeast genetics; yeast metabolism.
Copyright © 2022 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures


Similar articles
-
The Histone Deacetylases Hst1 and Rpd3 Integrate De Novo NAD+ Metabolism with Phosphate Sensing in Saccharomyces cerevisiae.Int J Mol Sci. 2023 Apr 28;24(9):8047. doi: 10.3390/ijms24098047. Int J Mol Sci. 2023. PMID: 37175754 Free PMC article.
-
The copper-sensing transcription factor Mac1, the histone deacetylase Hst1, and nicotinic acid regulate de novo NAD+ biosynthesis in budding yeast.J Biol Chem. 2019 Apr 5;294(14):5562-5575. doi: 10.1074/jbc.RA118.006987. Epub 2019 Feb 13. J Biol Chem. 2019. PMID: 30760525 Free PMC article.
-
Control of replication initiation by the Sum1/Rfm1/Hst1 histone deacetylase.BMC Mol Biol. 2008 Nov 6;9:100. doi: 10.1186/1471-2199-9-100. BMC Mol Biol. 2008. PMID: 18990212 Free PMC article.
-
Regulation of NAD+ metabolism, signaling and compartmentalization in the yeast Saccharomyces cerevisiae.DNA Repair (Amst). 2014 Nov;23:49-58. doi: 10.1016/j.dnarep.2014.07.009. Epub 2014 Aug 2. DNA Repair (Amst). 2014. PMID: 25096760 Free PMC article. Review.
-
Cross-talk in NAD+ metabolism: insights from Saccharomyces cerevisiae.Curr Genet. 2019 Oct;65(5):1113-1119. doi: 10.1007/s00294-019-00972-0. Epub 2019 Apr 16. Curr Genet. 2019. PMID: 30993413 Free PMC article. Review.
Cited by
-
The Histone Deacetylases Hst1 and Rpd3 Integrate De Novo NAD+ Metabolism with Phosphate Sensing in Saccharomyces cerevisiae.Int J Mol Sci. 2023 Apr 28;24(9):8047. doi: 10.3390/ijms24098047. Int J Mol Sci. 2023. PMID: 37175754 Free PMC article.
-
Metabolic profiling to evaluate the impact of amantadine and rimantadine on the secondary metabolism of a model organism.Sci Rep. 2023 Oct 5;13(1):16822. doi: 10.1038/s41598-023-43540-w. Sci Rep. 2023. PMID: 37798340 Free PMC article.
References
-
- Imai S., Armstrong C.M., Kaeberlein M., Guarente L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature. 2000;403:795–800. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

