High-yield genome engineering in primary cells using a hybrid ssDNA repair template and small-molecule cocktails
- PMID: 36008610
- PMCID: PMC10065198
- DOI: 10.1038/s41587-022-01418-8
High-yield genome engineering in primary cells using a hybrid ssDNA repair template and small-molecule cocktails
Abstract
Enhancing CRISPR-mediated site-specific transgene insertion efficiency by homology-directed repair (HDR) using high concentrations of double-stranded DNA (dsDNA) with Cas9 target sequences (CTSs) can be toxic to primary cells. Here, we develop single-stranded DNA (ssDNA) HDR templates (HDRTs) incorporating CTSs with reduced toxicity that boost knock-in efficiency and yield by an average of around two- to threefold relative to dsDNA CTSs. Using small-molecule combinations that enhance HDR, we could further increase knock-in efficiencies by an additional roughly two- to threefold on average. Our method works across a variety of target loci, knock-in constructs and primary human cell types, reaching HDR efficiencies of >80-90%. We demonstrate application of this approach for both pathogenic gene variant modeling and gene-replacement strategies for IL2RA and CTLA4 mutations associated with Mendelian disorders. Finally, we develop a good manufacturing practice (GMP)-compatible process for nonviral chimeric antigen receptor-T cell manufacturing, with knock-in efficiencies (46-62%) and yields (>1.5 × 109 modified cells) exceeding those of conventional approaches.
© 2022. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Competing interests
A.M. is a compensated cofounder, member of the boards of directors, and a member of the scientific advisory boards of Spotlight Therapeutics and Arsenal Biosciences. A.M. is a cofounder, member of the boards of directors, and a member of the scientific advisory board of Survey Genomics. A.M. is a compensated member of the scientific advisory board of NewLimit. A.M. owns stock in Arsenal Biosciences, Spotlight Therapeutics, NewLimit, Survey Genomics, PACT Pharma and Merck. A.M. has received fees from 23andMe, PACT Pharma, Juno Therapeutics, Trizell, Vertex, Merck, Amgen, Genentech, AlphaSights, Rupert Case Management, Bernstein and ALDA. A.M. is an investor in and informal advisor to Offline Ventures and a client of EPIQ. The Marson laboratory has received research support from Juno Therapeutics, Epinomics, Sanofi, GlaxoSmithKline, Gilead and Anthem. J.E. is a compensated cofounder at Mnemo Therapeutics. J.E. is a compensated scientific advisor to Cytovia Therapeutics. J.E. own stocks in Mnemo Therapeutica and Cytovia Therapeutics. J.E. has received a consulting fee from Casdin Capital. The Eyquem laboratory has received research support from Cytovia Therapeutic and Takeda. J.E. is a holder of patents pertaining to but not resulting from this work. H.L and L.Y. are employees of Genscript Biotech Corporation. J.W. has received consulting fees from Teneobio and Adaptive Biotech. D.N.N. receives consulting fees and sits on the scientific advisory board of Navan Technologies. T.L.R. is a cofounder, holds equity in, and is a member of the Scientific Advisory Board of Arsenal Bioscience. Discounted reagents were provided by Genscript. B.R.S., V.S.V. and A.M. are inventors on patent applications based on the findings described in this paper, a subset of which have been licensed by the University of California. A.H., A.T., W.G.P., Y.Y.C., F.B., E.S., S.V., M.R.M., J.J.C., R.Y., D.W., T.G.M., C.E.C. and J.H.E. declare no competing interests.
Figures













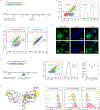

References
-
- Frangoul H et al. CRISPR-Cas9 gene editing for sickle cell disease and beta-thalassemia. New Engl. J. Med 384, 252–260 (2021). - PubMed
-
- US National Laboratory of Medicine. A safety and efficacy study evaluating ctx110 in subjects with relapsed or refractory B-cell malignancies (CARBON) ClinicalTrials.gov https://ClinicalTrials.gov/show/NCT04035434 (2019).
-
- US National Laboratory of Medicine. CRISPR-edited allogeneic anti-CD19 CAR-T cell therapy for relapsed/refractory B cell non-Hodgkin lymphoma ClinicalTrials.gov https://ClinicalTrials.gov/show/NCT04637763 (2020).
-
- US National Laboratory of Medicine. Transplantation of clustered regularly interspaced short palindromic repeats modified hematopoietic progenitor stem cells (CRISPR_SCD001) in patients with severe sickle cell disease ClinicalTrials.gov https://ClinicalTrials.gov/show/NCT04774536 (2021).
Publication types
MeSH terms
Substances
Grants and funding
- F30 DK120213/DK/NIDDK NIH HHS/United States
- K08 AI153767/AI/NIAID NIH HHS/United States
- T32 GM007618/GM/NIGMS NIH HHS/United States
- L40 AI140341/AI/NIAID NIH HHS/United States
- L30 TR002983/TR/NCATS NIH HHS/United States
- S10 OD025096/OD/NIH HHS/United States
- S10 RR028962/RR/NCRR NIH HHS/United States
- P01 AI155393/AI/NIAID NIH HHS/United States
- L30 AI140341/AI/NIAID NIH HHS/United States
- P01 AI138962/AI/NIAID NIH HHS/United States
- T32 AI007334/AI/NIAID NIH HHS/United States
- T32 DK007418/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

