Transcriptomic and Metabolomic Analysis of a Fusidic Acid-Selected fusA Mutant of Staphylococcus aureus
- PMID: 36009920
- PMCID: PMC9405211
- DOI: 10.3390/antibiotics11081051
Transcriptomic and Metabolomic Analysis of a Fusidic Acid-Selected fusA Mutant of Staphylococcus aureus
Abstract
Physiological experimentation, transcriptomics, and metabolomics were engaged to compare a fusidic acid-resistant Staphylococcus aureus mutant SH10001st-2 to its parent strain SH1000. SH10001st-2 harbored a mutation (H457Y) in the gene fusA which encodes the fusidic acid target, elongation factor G, as well as mutations in a putative phage gene of unknown function. SH10001st-2 grew slower than SH1000 at three temperatures and had reduced coagulase activity, two indicators of the fitness penalty reported for fusA-mediated fusidic acid- resistance in the absence of compensatory mutations. Despite the difference in growth rates, the levels of O2 consumption and CO2 production were comparable. Transcriptomic profiling revealed 326 genes were upregulated and 287 were downregulated in SH10001st-2 compared to SH1000. Cell envelope and transport and binding protein genes were the predominant functional categories of both upregulated and downregulated genes in SH10001st-2. Genes of virulence regulators, notably the agr and kdp systems, were highly upregulated as were genes encoding capsule production. Contrary to what is expected of mid-exponential phase cells, genes encoding secreted virulence factors were generally upregulated while those for adhesion-associated virulence factors were downregulated in SH10001st-2. Metabolomic analysis showed an overall increase in metabolite pools in SH10001st-2 compared to SH1000, mostly for amino acids and sugars. Slowed growth and metabolite accumulation may be byproducts of fusA mutation-mediated protein synthesis impairment, but the overall results indicate that SH10001st-2 is compensating for the H457Y fitness penalty by repurposing its virulence machinery, in conjunction with increasing metabolite uptake capacity, in order to increase nutrient acquisition.
Keywords: Staphylococcus aureus; fusidic acid-resistance; metabolomics; respiration; transcriptional profiling.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Molecular analysis of fusidic acid resistance in Staphylococcus aureus.Mol Microbiol. 2003 Jan;47(2):463-9. doi: 10.1046/j.1365-2958.2003.03307.x. Mol Microbiol. 2003. PMID: 12519196
-
Compensatory adaptation to the loss of biological fitness associated with acquisition of fusidic acid resistance in Staphylococcus aureus.Antimicrob Agents Chemother. 2005 Apr;49(4):1426-31. doi: 10.1128/AAC.49.4.1426-1431.2005. Antimicrob Agents Chemother. 2005. PMID: 15793122 Free PMC article.
-
The fusidic acid stimulon of Staphylococcus aureus.J Antimicrob Chemother. 2008 Dec;62(6):1207-14. doi: 10.1093/jac/dkn363. Epub 2008 Sep 10. J Antimicrob Chemother. 2008. PMID: 18786940 Free PMC article.
-
Dumb and dumber--the potential waste of a useful antistaphylococcal agent: emerging fusidic acid resistance in Staphylococcus aureus.Clin Infect Dis. 2006 Feb 1;42(3):394-400. doi: 10.1086/499365. Epub 2005 Dec 15. Clin Infect Dis. 2006. PMID: 16392088 Review.
-
Characterization of global patterns and the genetics of fusidic acid resistance.Clin Infect Dis. 2011 Jun;52 Suppl 7:S487-92. doi: 10.1093/cid/cir164. Clin Infect Dis. 2011. PMID: 21546625 Review.
Cited by
-
Distribution of Antimicrobial Resistance and Biofilm Production Genes in the Genomic Sequences of S. aureus: A Global In Silico Analysis.Antibiotics (Basel). 2025 Apr 1;14(4):364. doi: 10.3390/antibiotics14040364. Antibiotics (Basel). 2025. PMID: 40298499 Free PMC article.
-
Enterococcus faecalis OG1RF Evolution at Low pH Selects Fusidate-Sensitive Mutants in Elongation Factor G and at High pH Selects Defects in Phosphate Transport.Appl Environ Microbiol. 2023 Jun 28;89(6):e0046623. doi: 10.1128/aem.00466-23. Epub 2023 Jun 5. Appl Environ Microbiol. 2023. PMID: 37272807 Free PMC article.
-
Clinical Features and Antibiotic Susceptibility of Staphylococcus aureus-Infected Dermatoses.J Clin Med. 2025 Feb 8;14(4):1084. doi: 10.3390/jcm14041084. J Clin Med. 2025. PMID: 40004615 Free PMC article.
-
Biochemical and Cellular Characterization of the Function of Fluorophosphonate-Binding Hydrolase H (FphH) in Staphylococcus aureus Support a Role in Bacterial Stress Response.ACS Infect Dis. 2023 Nov 10;9(11):2119-2132. doi: 10.1021/acsinfecdis.3c00246. Epub 2023 Oct 12. ACS Infect Dis. 2023. PMID: 37824340 Free PMC article.
-
Vancomycin-variable enterococci in sheep and cattle isolates and whole-genome sequencing analysis of isolates harboring vanM and vanB genes.Iran J Vet Res. 2023;24(3):182-192. doi: 10.22099/IJVR.2023.47465.6855. Iran J Vet Res. 2023. PMID: 38269016 Free PMC article.
References
-
- Kourtis A.P., Hatfield K., Baggs J., Mu Y., See I., Epson E., Nadle J., Kainer M.A., Dumyati G., Petit S., et al. Vital Signs: Epidemiology and Recent Trends in Methicillin-Resistant and in Methicillin-Susceptible Staphylococcus aureus Bloodstream Infections—United States. Morb. Mortal. Wkly. Rep. 2019;68:214–219. doi: 10.15585/mmwr.mm6809e1. - DOI - PMC - PubMed
-
- Hajikhani B., Goudarzi M., Kakavandi S., Amini S., Zamani S., van Belkum A., Goudarzi H., Dadashi M. The global prevalence of fusidic acid resistance in clinical isolates of Staphylococcus aureus: A systematic review and meta-analysis. Antimicrob. Resist. Infect. Control. 2021;10:75. doi: 10.1186/s13756-021-00943-6. - DOI - PMC - PubMed
-
- Guo X., Peisker K., Backbro K., Chen Y., Koripella R.K., Mandava C.S., Sanyal S., Selmer M. Structure and function of FusB: An elongation factor G-binding fusidic acid resistance protein active in ribosomal translocation and recycling. Open Biol. 2012;2:120016. doi: 10.1098/rsob.120016. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

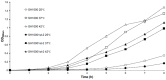
 ,
,  ) 37 °C (
) 37 °C ( ,
,  ), or 42 °C (
), or 42 °C ( ,
,  ). All data shown are the mean of triplicate experiments and error bars represent the standard deviation.
). All data shown are the mean of triplicate experiments and error bars represent the standard deviation.