Genomic Analyses of Non-Coding RNAs Overlapping Transposable Elements and Its Implication to Human Diseases
- PMID: 36012216
- PMCID: PMC9409130
- DOI: 10.3390/ijms23168950
Genomic Analyses of Non-Coding RNAs Overlapping Transposable Elements and Its Implication to Human Diseases
Abstract
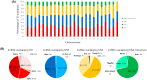
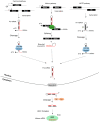
It is estimated that up to 80% of the human genome is transcribed into RNA molecules but less than 2% of the genome encodes the proteins, and the rest of the RNA transcripts that are not translated into protein are called non-coding RNAs (ncRNAs). Many studies have revealed that ncRNAs have biochemical activities as epigenetic regulators at the post-transcriptional level. Growing evidence has demonstrated that transposable elements (TEs) contribute to a large percentage of ncRNAs' transcription. The TEs inserted into certain parts of the genome can act as alternative promoters, enhancers, and insulators, and the accumulation of TEs increases genetic diversity in the human genome. The TEs can also generate microRNAs, so-called miRNA-derived from transposable elements (MDTEs), and are also implicated in disease progression, such as infectious diseases and cancer. Here, we analyzed the origin of ncRNAs and reviewed the published literature on MDTEs related to disease progression.
Keywords: MDTE; disease; long non-coding RNA; non-coding RNA; transposable element.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

