Application of Neuromelanin MR Imaging in Parkinson Disease
- PMID: 36017746
- PMCID: PMC10086789
- DOI: 10.1002/jmri.28414
Application of Neuromelanin MR Imaging in Parkinson Disease
Abstract
MRI has been used to develop biomarkers for movement disorders such as Parkinson disease (PD) and other neurodegenerative disorders with parkinsonism such as progressive supranuclear palsy and multiple system atrophy. One of these imaging biomarkers is neuromelanin (NM), whose integrity can be assessed from its contrast and volume. NM is found mainly in certain brain stem structures, namely, the substantia nigra pars compacta (SNpc), the ventral tegmental area, and the locus coeruleus. Another major biomarker is brain iron, which often increases in concert with NM degeneration. These biomarkers have the potential to improve diagnostic certainty in differentiating between PD and other neurodegenerative disorders similar to PD, as well as provide a better understanding of pathophysiology. Mapping NM in vivo has clinical importance for gauging the premotor phase of PD when there is a greater than 50% loss of dopaminergic SNpc melanized neurons. As a metal ion chelator, NM can absorb iron. When NM is released from neurons, it deposits iron into the intracellular tissues of the SNpc; the result is iron that can be imaged and measured using quantitative susceptibility mapping. An increase of iron also leads to the disappearance of the nigrosome-1 sign, another neuroimage biomarker for PD. Therefore, mapping NM and iron changes in the SNpc are a practical means for improving early diagnosis of PD and in monitoring disease progression. In this review, we discuss the functions and location of NM, how NM-MRI is performed, the automatic mapping of NM and iron content, how NM-related imaging biomarkers can be used to enhance PD diagnosis and differentiate it from other neurodegenerative disorders, and potential advances in NM imaging methods. With major advances currently evolving for rapid imaging and artificial intelligence, NM-related biomarkers are likely to have increasingly important roles for enhancing diagnostic capabilities in PD. EVIDENCE LEVEL: 1 TECHNICAL EFFICACY: Stage 2.
Keywords: Parkinson disease; magnetic resonance imaging; magnetization transfer contrast; neuromelanin; nigrosome 1.
© 2022 The Authors. Journal of Magnetic Resonance Imaging published by Wiley Periodicals LLC on behalf of International Society for Magnetic Resonance in Medicine.
Figures
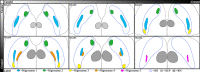

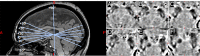

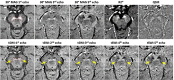
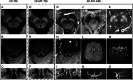
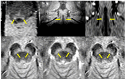

References
-
- Fearnley JM, Lees AJ. Ageing and Parkinson's disease: Substantia nigra regional selectivity. Brain 1991;114(Pt 5):2283‐2301. - PubMed
-
- Damier P, Hirsch E, Agid Y, Graybiel A. The substantia nigra of the human brain: II. Patterns of loss of dopamine‐containing neurons in Parkinson's disease. Brain 1999;122(8):1437‐1448. - PubMed
-
- Wypijewska A, Galazka‐Friedman J, Bauminger ER, et al. Iron and reactive oxygen species activity in parkinsonian substantia nigra. Parkinsonism Relat Disord 2010;16(5):329‐333. - PubMed
-
- He N, Ghassaban K, Huang P, et al. Imaging iron and neuromelanin simultaneously using a single 3D gradient echo magnetization transfer sequence: Combining neuromelanin, iron and the nigrosome‐1 sign as complementary imaging biomarkers in early stage Parkinson's disease. Neuroimage 2021;230:117810. - PubMed
-
- Takahashi H, Watanabe Y, Tanaka H, et al. Comprehensive MRI quantification of the substantia nigra pars compacta in Parkinson's disease. Eur J Radiol 2018;109:48‐56. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

