Additive genetic effects in interacting species jointly determine the outcome of caterpillar herbivory
- PMID: 36037349
- PMCID: PMC9456756
- DOI: 10.1073/pnas.2206052119
Additive genetic effects in interacting species jointly determine the outcome of caterpillar herbivory
Abstract
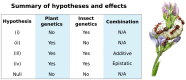
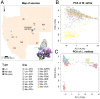
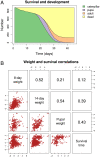
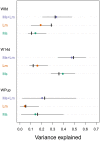
Plant-insect interactions are common and important in basic and applied biology. Trait and genetic variation can affect the outcome and evolution of these interactions, but the relative contributions of plant and insect genetic variation and how these interact remain unclear and are rarely subject to assessment in the same experimental context. Here, we address this knowledge gap using a recent host-range expansion onto alfalfa by the Melissa blue butterfly. Common garden rearing experiments and genomic data show that caterpillar performance depends on plant and insect genetic variation, with insect genetics contributing to performance earlier in development and plant genetics later. Our models of performance based on caterpillar genetics retained predictive power when applied to a second common garden. Much of the plant genetic effect could be explained by heritable variation in plant phytochemicals, especially saponins, peptides, and phosphatidyl cholines, providing a possible mechanistic understanding of variation in the species interaction. We find evidence of polygenic, mostly additive effects within and between species, with consistent effects of plant genotype on growth and development across multiple butterfly species. Our results inform theories of plant-insect coevolution and the evolution of diet breadth in herbivorous insects and other host-specific parasites.
Keywords: coevolution; genomic prediction; phytochemicals; plant–insect interaction; polygenic.
Conflict of interest statement
The authors declare no competing interest.
Figures

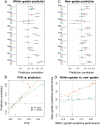

References
-
- Manceau M., Domingues V. S., Mallarino R., Hoekstra H. E., The developmental role of Agouti in color pattern evolution. Science 331, 1062–1065 (2011). - PubMed
-
- Van’t Hof A. E., et al. ., The industrial melanism mutation in British peppered moths is a transposable element. Nature 534, 102–105 (2016). - PubMed
-
- Villoutreix R., et al. ., Large-scale mutation in the evolution of a gene complex for cryptic coloration. Science 369, 460–466 (2020). - PubMed
-
- Nylin S., et al. ., Embracing colonizations: A new paradigm for species association dynamics. Trends Ecol. Evol. 33, 4–14 (2018). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

