TTYH family members form tetrameric complexes at the cell membrane
- PMID: 36042377
- PMCID: PMC9427776
- DOI: 10.1038/s42003-022-03862-3
TTYH family members form tetrameric complexes at the cell membrane
Abstract
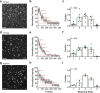
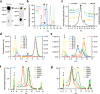
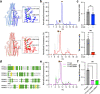
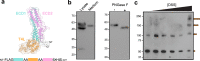
The conserved Tweety homolog (TTYH) family consists of three paralogs in vertebrates, displaying a ubiquitous expression pattern. Although considered as ion channels for almost two decades, recent structural and functional analyses refuted this role. Intriguingly, while all paralogs shared a dimeric stoichiometry following detergent solubilization, their structures revealed divergence in their relative subunit orientation. Here, we determined the stoichiometry of intact mouse TTYH (mTTYH) complexes in cells. Using cross-linking and single-molecule fluorescence microscopy, we demonstrate that mTTYH1 and mTTYH3 form tetramers at the plasma membrane, stabilized by interactions between their extracellular domains. Using blue-native PAGE, fluorescence-detection size-exclusion chromatography, and hydrogen/deuterium exchange mass spectrometry (HDX-MS), we reveal that detergent solubilization results in tetramers destabilization, leading to their dissolution into dimers. Moreover, HDX-MS demonstrates that the extracellular domains are stabilized in the context of the tetrameric mTTYH complex. Together, our results expose the innate tetrameric organization of TTYH complexes at the cell membrane. Future structural analyses of these assemblies in native membranes are required to illuminate their long-sought cellular function.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


References
-
- Campbell HD, et al. The Drosophila melanogaster flightless-I gene involved in gastrulation and muscle degeneration encodes gelsolin-like and leucine-rich repeat domains and is conserved in Caenorhabditis elegans and humans. Proc. Natl Acad. Sci. USA. 1993;90:11386–11390. doi: 10.1073/pnas.90.23.11386. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

