Dynamic genome evolution in a model fern
- PMID: 36050461
- PMCID: PMC9477723
- DOI: 10.1038/s41477-022-01226-7
Dynamic genome evolution in a model fern
Abstract
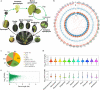
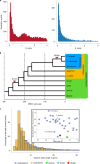
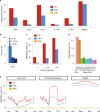
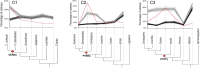
The large size and complexity of most fern genomes have hampered efforts to elucidate fundamental aspects of fern biology and land plant evolution through genome-enabled research. Here we present a chromosomal genome assembly and associated methylome, transcriptome and metabolome analyses for the model fern species Ceratopteris richardii. The assembly reveals a history of remarkably dynamic genome evolution including rapid changes in genome content and structure following the most recent whole-genome duplication approximately 60 million years ago. These changes include massive gene loss, rampant tandem duplications and multiple horizontal gene transfers from bacteria, contributing to the diversification of defence-related gene families. The insertion of transposable elements into introns has led to the large size of the Ceratopteris genome and to exceptionally long genes relative to other plants. Gene family analyses indicate that genes directing seed development were co-opted from those controlling the development of fern sporangia, providing insights into seed plant evolution. Our findings and annotated genome assembly extend the utility of Ceratopteris as a model for investigating and teaching plant biology.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
-
- Kenrick P, Crane PR. The origin and early evolution of plants on land. Nature. 1997;389:33–39. doi: 10.1038/37918. - DOI
-
- Lloyd RM. Mating systems and genetic load in pioneer and non-pioneer Hawaiian Pteridophyta. Bot. J. Linn. Soc. 1974;69:23–35. doi: 10.1111/j.1095-8339.1974.tb01611.x. - DOI
-
- de León SG, Briones O, Aguirre A, Mehltreter K, Pérez-García B. Germination of an invasive fern responds better than native ferns to water and light stress in a Mexican cloud forest. Biol. Invas. 2021;20:3187–3199. doi: 10.1007/s10530-021-02570-z. - DOI
-
- Raja W, Rathaur P, John SA, Ramteke PW. Azolla: an aquatic pteridophyte with great potential. Int. J. Res. Biol. Sci. 2012;2:68–72.

