The Four Causes: The Functional Architecture of Centromeres and Kinetochores
- PMID: 36055650
- PMCID: PMC7614391
- DOI: 10.1146/annurev-genet-072820-034559
The Four Causes: The Functional Architecture of Centromeres and Kinetochores
Abstract
Kinetochores are molecular machines that power chromosome segregation during the mitotic and meiotic cell divisions of all eukaryotes. Aristotle explains how we think we have knowledge of a thing only when we have grasped its cause. In our case, to gain understanding of the kinetochore, the four causes correspond to questions that we must ask: (a) What are the constituent parts, (b) how does it assemble, (c) what is the structure and arrangement, and (d) what is the function? Here we outline the current blueprint for the assembly of a kinetochore, how functions are mapped onto this architecture, and how this is shaped by the underlying pericentromeric chromatin. The view of the kinetochore that we present is possible because an almost complete parts list of the kinetochore is now available alongside recent advances using in vitro reconstitution, structural biology, and genomics. In many organisms, each kinetochore binds to multiple microtubules, and we propose a model for how this ensemble-level architecture is organized, drawing on key insights from the simple one microtubule-one kinetochore setup in budding yeast and innovations that enable meiotic chromosome segregation.
Keywords: cell division; centromere; chromatin; chromosome; kinetochore; meiosis; microtubule; mitosis; mitotic spindle.
Figures
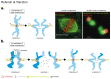
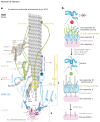
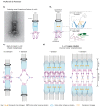
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

