Identification of novel neutralizing determinants for protection against HCV
- PMID: 36056620
- PMCID: PMC9936975
- DOI: 10.1002/hep.32772
Identification of novel neutralizing determinants for protection against HCV
Abstract
Background and aims: HCV evasion of neutralizing antibodies (nAb) results in viral persistence and poses challenges to the development of an urgently needed vaccine. N-linked glycosylation of viral envelope proteins is a key mechanism for such evasion. To facilitate rational vaccine design, we aimed to identify determinants of protection of conserved neutralizing epitopes.
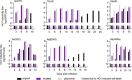
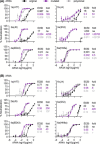
Approach and results: Using a reverse evolutionary approach, we passaged genotype 1a, 1b, 2a, 3a, and 4a HCV with envelope proteins (E1 and E2) derived from chronically infected patients without selective pressure by nAb in cell culture. Compared with the original viruses, HCV recombinants, engineered to harbor substitutions identified in polyclonal cell culture-passaged viruses, showed highly increased fitness and exposure of conserved neutralizing epitopes in antigenic regions 3 and 4, associated with protection from chronic infection. Further reverse genetic studies of acquired E1/E2 substitutions identified positions 418 and 532 in the N1 and N6 glycosylation motifs, localizing to adjacent E2 areas, as key regulators of changes of the E1/E2 conformational state, which governed viral sensitivity to nAb. These effects were independent of predicted glycan occupancy.
Conclusions: We show how N-linked glycosylation motifs can trigger dramatic changes in HCV sensitivity to nAb, independent of glycan occupancy. These findings aid in the understanding of HCV nAb evasion and rational vaccine design, as they can be exploited to stabilize the structurally flexible envelope proteins in an open conformation, exposing important neutralizing epitopes. Finally, this work resulted in a panel of highly fit cell culture infectious HCV recombinants.
Copyright © 2023 The Author(s). Published by Wolters Kluwer Health, Inc. on behalf of American Association for the Study of Liver Diseases.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




References
-
- Simmonds P. The origin of hepatitis C virus. Curr Top Microbiol Immunol. 2013;369:1–15. - PubMed
-
- Cox AL. Global control of hepatitis C virus. 2015;349:790–1. - PubMed
-
- Bartenschlager R, Baumert TF, Bukh J, Houghton M, Lemon SM, Lindenbach BD, et al. . Critical challenges and emerging opportunities in hepatitis C virus research in an era of potent antiviral therapy: considerations for scientists and funding agencies. Virus Res. 2018;248:53–62. - PubMed
-
- Bukh J. The history of hepatitis C virus (HCV): basic research reveals unique features in phylogeny, evolution and the viral life cycle with new perspectives for epidemic control. J Hepatol. 2016;65:S2–21. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

