Micronuclei from misaligned chromosomes that satisfy the spindle assembly checkpoint in cancer cells
- PMID: 36057259
- PMCID: PMC9559752
- DOI: 10.1016/j.cub.2022.08.026
Micronuclei from misaligned chromosomes that satisfy the spindle assembly checkpoint in cancer cells
Abstract
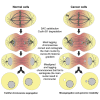
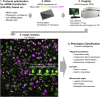
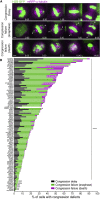
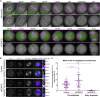
Chromosome alignment to the spindle equator is a hallmark of mitosis thought to promote chromosome segregation fidelity in metazoans. Yet chromosome alignment is only indirectly supervised by the spindle assembly checkpoint (SAC) as a byproduct of chromosome bi-orientation, and the consequences of defective chromosome alignment remain unclear. Here, we investigated how human cells respond to chromosome alignment defects of distinct molecular nature by following the fate of live HeLa cells after RNAi-mediated depletion of 125 proteins previously implicated in chromosome alignment. We confirmed chromosome alignment defects upon depletion of 108/125 proteins. Surprisingly, in all confirmed cases, depleted cells frequently entered anaphase after a delay with misaligned chromosomes. Using depletion of prototype proteins resulting in defective chromosome alignment, we show that misaligned chromosomes often satisfy the SAC and directly missegregate without lagging behind in anaphase. In-depth analysis of specific molecular perturbations that prevent proper kinetochore-microtubule attachments revealed that misaligned chromosomes that missegregate frequently result in micronuclei. Higher-resolution live-cell imaging indicated that, contrary to most anaphase lagging chromosomes that correct and reintegrate the main nuclei, misaligned chromosomes are a strong predictor of micronuclei formation in a cancer cell model of chromosomal instability, but not in non-transformed near-diploid cells. We provide evidence supporting that intrinsic differences in kinetochore-microtubule attachment stability on misaligned chromosomes account for this distinct outcome. Thus, misaligned chromosomes that satisfy the SAC may represent a previously overlooked mechanism driving chromosomal/genomic instability during cancer cell division, and we unveil genetic conditions predisposing for these events.
Keywords: Mad2; aneuploidy; cancer; chromosomal instability; chromosome congression; cyclin B1; kinetochore; micronuclei; mitosis; spindle assembly checkpoint.
Copyright © 2022 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests B.O. declares that he is a consultant specialist at Volastra Therapeutics.
Figures




Similar articles
-
Double-checking chromosome segregation.J Cell Biol. 2023 May 1;222(5):e202301106. doi: 10.1083/jcb.202301106. Epub 2023 Apr 5. J Cell Biol. 2023. PMID: 37017932 Free PMC article. Review.
-
NuMA deficiency causes micronuclei via checkpoint-insensitive k-fiber minus-end detachment from mitotic spindle poles.Curr Biol. 2023 Feb 6;33(3):572-580.e2. doi: 10.1016/j.cub.2022.12.017. Epub 2023 Jan 9. Curr Biol. 2023. PMID: 36626904
-
Phosphorylation of the Ndc80 complex protein, HEC1, by Nek2 kinase modulates chromosome alignment and signaling of the spindle assembly checkpoint.Mol Biol Cell. 2011 Oct;22(19):3584-94. doi: 10.1091/mbc.E11-01-0012. Epub 2011 Aug 10. Mol Biol Cell. 2011. PMID: 21832156 Free PMC article.
-
Chromosome Segregation Is Biased by Kinetochore Size.Curr Biol. 2018 May 7;28(9):1344-1356.e5. doi: 10.1016/j.cub.2018.03.023. Epub 2018 Apr 26. Curr Biol. 2018. PMID: 29706521 Free PMC article.
-
Connecting up and clearing out: how kinetochore attachment silences the spindle assembly checkpoint.Chromosoma. 2012 Oct;121(5):509-25. doi: 10.1007/s00412-012-0378-5. Epub 2012 Jul 11. Chromosoma. 2012. PMID: 22782189 Review.
Cited by
-
Cul3 substrate adaptor SPOP targets Nup153 for degradation.Mol Biol Cell. 2025 Mar 1;36(3):ar24. doi: 10.1091/mbc.E24-04-0198. Epub 2025 Jan 9. Mol Biol Cell. 2025. PMID: 39785820 Free PMC article.
-
Mitotic DNA damage promotes chromokinesin-mediated missegregation of polar chromosomes in cancer cells.Mol Biol Cell. 2023 May 1;34(5):ar47. doi: 10.1091/mbc.E22-11-0518. Epub 2023 Mar 29. Mol Biol Cell. 2023. PMID: 36989031 Free PMC article.
-
Study of genotoxic and cytotoxic effects induced in human fibroblasts by exposure to pulsed and continuous 1.6 GHz radiofrequency.Front Public Health. 2024 Jul 31;12:1419525. doi: 10.3389/fpubh.2024.1419525. eCollection 2024. Front Public Health. 2024. PMID: 39145180 Free PMC article.
-
Double-checking chromosome segregation.J Cell Biol. 2023 May 1;222(5):e202301106. doi: 10.1083/jcb.202301106. Epub 2023 Apr 5. J Cell Biol. 2023. PMID: 37017932 Free PMC article. Review.
-
A scoping review and evidence map of radiofrequency field exposure and genotoxicity: assessing in vivo, in vitro, and epidemiological data.Front Public Health. 2025 Jul 30;13:1613353. doi: 10.3389/fpubh.2025.1613353. eCollection 2025. Front Public Health. 2025. PMID: 40809778 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

