From descriptive connectome to mechanistic connectome: Generative modeling in functional magnetic resonance imaging analysis
- PMID: 36061504
- PMCID: PMC9428697
- DOI: 10.3389/fnhum.2022.940842
From descriptive connectome to mechanistic connectome: Generative modeling in functional magnetic resonance imaging analysis
Abstract
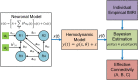
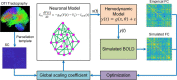
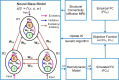
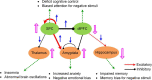
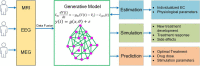
As a newly emerging field, connectomics has greatly advanced our understanding of the wiring diagram and organizational features of the human brain. Generative modeling-based connectome analysis, in particular, plays a vital role in deciphering the neural mechanisms of cognitive functions in health and dysfunction in diseases. Here we review the foundation and development of major generative modeling approaches for functional magnetic resonance imaging (fMRI) and survey their applications to cognitive or clinical neuroscience problems. We argue that conventional structural and functional connectivity (FC) analysis alone is not sufficient to reveal the complex circuit interactions underlying observed neuroimaging data and should be supplemented with generative modeling-based effective connectivity and simulation, a fruitful practice that we term "mechanistic connectome." The transformation from descriptive connectome to mechanistic connectome will open up promising avenues to gain mechanistic insights into the delicate operating principles of the human brain and their potential impairments in diseases, which facilitates the development of effective personalized treatments to curb neurological and psychiatric disorders.
Keywords: Dynamic Neural Model; biophysical network model; brain network; computational modeling; connectome; dynamical causal model; fMRI; neural mass model.
Copyright © 2022 Li and Yap.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Structural Basis of Large-Scale Functional Connectivity in the Mouse.J Neurosci. 2017 Aug 23;37(34):8092-8101. doi: 10.1523/JNEUROSCI.0438-17.2017. Epub 2017 Jul 17. J Neurosci. 2017. PMID: 28716961 Free PMC article.
-
Whole-brain estimates of directed connectivity for human connectomics.Neuroimage. 2021 Jan 15;225:117491. doi: 10.1016/j.neuroimage.2020.117491. Epub 2020 Oct 24. Neuroimage. 2021. PMID: 33115664
-
A generative model of the connectome with dynamic axon growth.Netw Neurosci. 2024 Dec 10;8(4):1192-1211. doi: 10.1162/netn_a_00397. eCollection 2024. Netw Neurosci. 2024. PMID: 39735503 Free PMC article.
-
Biophysical Modeling of Large-Scale Brain Dynamics and Applications for Computational Psychiatry.Biol Psychiatry Cogn Neurosci Neuroimaging. 2018 Sep;3(9):777-787. doi: 10.1016/j.bpsc.2018.07.004. Epub 2018 Jul 19. Biol Psychiatry Cogn Neurosci Neuroimaging. 2018. PMID: 30093344 Free PMC article. Review.
-
Great expectations: using whole-brain computational connectomics for understanding neuropsychiatric disorders.Neuron. 2014 Dec 3;84(5):892-905. doi: 10.1016/j.neuron.2014.08.034. Neuron. 2014. PMID: 25475184 Review.
Cited by
-
Causation in neuroscience: keeping mechanism meaningful.Nat Rev Neurosci. 2024 Feb;25(2):81-90. doi: 10.1038/s41583-023-00778-7. Epub 2024 Jan 11. Nat Rev Neurosci. 2024. PMID: 38212413 Review.
-
Recent Progress in Brain Network Models for Medical Applications: A Review.Health Data Sci. 2024 Jul 8;4:0157. doi: 10.34133/hds.0157. eCollection 2024. Health Data Sci. 2024. PMID: 38979037 Free PMC article. Review.
-
Development of Effective Connectome from Infancy to Adolescence.Med Image Comput Comput Assist Interv. 2024 Oct;15003:131-140. doi: 10.1007/978-3-031-72384-1_13. Epub 2024 Oct 3. Med Image Comput Comput Assist Interv. 2024. PMID: 39866850
-
A novel brain tumor magnetic resonance imaging dataset (Gazi Brains 2020): initial benchmark results and comprehensive analysis.PeerJ Comput Sci. 2025 Jun 10;11:e2920. doi: 10.7717/peerj-cs.2920. eCollection 2025. PeerJ Comput Sci. 2025. PMID: 40567687 Free PMC article.
-
Macroscale coupling between structural and effective connectivity in the mouse brain.Sci Rep. 2024 Feb 7;14(1):3142. doi: 10.1038/s41598-024-51613-7. Sci Rep. 2024. PMID: 38326324 Free PMC article.
References
-
- Ashby F. G., Spiering B. J. (2004). The neurobiology of category learning. Behav. Cogn. Neurosci. Rev. 3 101–113. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

