Transient proteolysis reduction of Nicotiana benthamiana-produced CAP256 broadly neutralizing antibodies using CRISPR/Cas9
- PMID: 36061808
- PMCID: PMC9433777
- DOI: 10.3389/fpls.2022.953654
Transient proteolysis reduction of Nicotiana benthamiana-produced CAP256 broadly neutralizing antibodies using CRISPR/Cas9
Erratum in
-
Corrigendum: Transient proteolysis reduction of Nicotiana benthamiana-produced CAP256 broadly neutralizing antibodies using CRISPR/Cas9.Front Plant Sci. 2025 Mar 19;16:1568756. doi: 10.3389/fpls.2025.1568756. eCollection 2025. Front Plant Sci. 2025. PMID: 40177014 Free PMC article.
Abstract
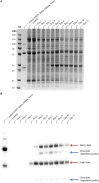
The hypersensitive response is elicited by Agrobacterium infiltration of Nicotiana benthamiana, including the induction and accumulation of pathogenesis-related proteins, such as proteases. This includes the induction of the expression of several cysteine proteases from the C1 (papain-like cysteine protease) and C13 (legumain-like cysteine protease) families. This study demonstrates the role of cysteine proteases: NbVPE-1a, NbVPE-1b, and NbCysP6 in the proteolytic degradation of Nicotiana benthamiana (glycosylation mutant ΔXTFT)-produced anti-human immunodeficiency virus broadly neutralizing antibody, CAP256-VRC26.25. Three putative cysteine protease cleavage sites were identified in the fragment crystallizable region. We further demonstrate the transient coexpression of CAP256-VRC26.25 with CRISPR/Cas9-mediated genome editing vectors targeting the NbVPE-1a, NbVPE-1b, and NbCysP6 genes which resulted in a decrease in CAP256-VRC26.25 degradation. No differences in structural features were observed between the human embryonic kidney 293 (HEK293)-produced and ΔXTFT broadly neutralizing antibodies produced with and without the coexpression of genome-editing vectors. Furthermore, despite the presence of proteolytically degraded fragments of plant-produced CAP256-VRC26.25 without the coexpression of genome editing vectors, no influence on the in vitro functional activity was detected. Collectively, we demonstrate an innovative in planta strategy for improving the quality of the CAP256 antibodies through the transient expression of the CRISPR/Cas9 vectors.
Keywords: CRISPR/Cas9; Nicotiana benthamiana; genome editing; human immunodeficiency virus; immunoglobulin G; plant biotechnology; proteases.
Copyright © 2022 Singh, Pillay, Naicker, Alexandre, Malatji, Mach, Steinkellner, Vorster, Chikwamba and Tsekoa.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Corrigendum: Transient proteolysis reduction of Nicotiana benthamiana-produced CAP256 broadly neutralizing antibodies using CRISPR/Cas9.Front Plant Sci. 2025 Mar 19;16:1568756. doi: 10.3389/fpls.2025.1568756. eCollection 2025. Front Plant Sci. 2025. PMID: 40177014 Free PMC article.
-
A plant-biotechnology approach for producing highly potent anti-HIV antibodies for antiretroviral therapy consideration.J Genet Eng Biotechnol. 2021 Dec 8;19(1):180. doi: 10.1186/s43141-021-00279-z. J Genet Eng Biotechnol. 2021. PMID: 34878628 Free PMC article.
-
The papain-like cysteine proteinases NbCysP6 and NbCysP7 are highly processive enzymes with substrate specificities complementary to Nicotiana benthamiana cathepsin B.Biochim Biophys Acta Proteins Proteom. 2017 Apr;1865(4):444-452. doi: 10.1016/j.bbapap.2017.02.007. Epub 2017 Feb 7. Biochim Biophys Acta Proteins Proteom. 2017. PMID: 28188928
-
The Enhancement of Plant Disease Resistance Using CRISPR/Cas9 Technology.Front Plant Sci. 2018 Aug 24;9:1245. doi: 10.3389/fpls.2018.01245. eCollection 2018. Front Plant Sci. 2018. PMID: 30197654 Free PMC article. Review.
-
CRISPR/Cas9 genome editing through in planta transformation.Crit Rev Biotechnol. 2020 Mar;40(2):153-168. doi: 10.1080/07388551.2019.1709795. Epub 2020 Jan 5. Crit Rev Biotechnol. 2020. PMID: 31903793 Review.
Cited by
-
Harnessing Transient Expression Systems with Plant Viral Vectors for the Production of Biopharmaceuticals in Nicotiana benthamiana.Int J Mol Sci. 2025 Jun 9;26(12):5510. doi: 10.3390/ijms26125510. Int J Mol Sci. 2025. PMID: 40564973 Free PMC article. Review.
-
Unlocking the full potential of plant cell-based production for valuable proteins: Challenges and innovative strategies.Biotechnol Adv. 2025 Mar-Apr;79:108526. doi: 10.1016/j.biotechadv.2025.108526. Epub 2025 Feb 4. Biotechnol Adv. 2025. PMID: 39914685 Review.
-
Crop genome editing through tissue-culture-independent transformation methods.Front Genome Ed. 2024 Dec 5;6:1490295. doi: 10.3389/fgeed.2024.1490295. eCollection 2024. Front Genome Ed. 2024. PMID: 39703881 Free PMC article. Review.
-
Improving transient protein expression in agroinfiltrated Nicotiana benthamiana.New Phytol. 2024 Aug;243(3):846-850. doi: 10.1111/nph.19894. Epub 2024 Jun 7. New Phytol. 2024. PMID: 38849321 Free PMC article. Review.
References
-
- Brezski R. J., Vafa O., Petrone D., Tam S. H., Powers G., Ryan M. H., et al. . (2009). Tumor-associated and microbial proteases compromise host IgG effector functions by a single cleavage proximal to the hinge. Proc. Natl. Acad. Sci. U. S. A. 106, 17864–17869. 10.1073/pnas.0904174106 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

