Development of a panel of three multiplex allele-specific qRT-PCR assays for quick differentiation of recombinant variants and Omicron subvariants of SARS-CoV-2
- PMID: 36061868
- PMCID: PMC9433905
- DOI: 10.3389/fcimb.2022.953027
Development of a panel of three multiplex allele-specific qRT-PCR assays for quick differentiation of recombinant variants and Omicron subvariants of SARS-CoV-2
Abstract
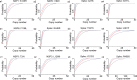
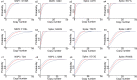
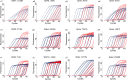
Quick differentiation of the circulating variants and the emerging recombinant variants of SARS-CoV-2 is essential to monitor their transmission. However, the widely used gene sequencing method is time-consuming and costly when facing the viral recombinant variants, because partial or whole genome sequencing is required. Allele-specific real time RT-PCR (qRT-PCR) represents a quick and cost-effective method in SNP genotyping and has been successfully applied for SARS-CoV-2 variant screening. In the present study, we developed a panel of 3 multiplex allele-specific qRT-PCR assays targeting 12 key differential mutations for quick differentiation of SARS-CoV-2 recombinant variants (XD and XE) and Omicron subvariants (BA.1 and BA.2). Two parallel multiplex qRT-PCR reactions were designed to separately target the protype allele and the mutated allele of the four mutations in each allele-specific qRT-PCR assay. The variation of Cp values (ΔCp) between the two multiplex qRT-PCR reactions was applied for mutation determination. The developed multiplex allele-specific qRT-PCR assays exhibited outstanding analytical sensitivities (with limits of detection [LoDs] of 2.97-27.43 copies per reaction), wide linear detection ranges (107-100 copies per reaction), good amplification efficiencies (82% to 95%), good reproducibility (Coefficient of Variations (CVs) < 5% in both intra-assay and inter-assay tests) and clinical performances (99.5%-100% consistency with Sanger sequencing). The developed multiplex allele-specific qRT-PCR assays in this study provide an alternative tool for quick differentiation of SARS-CoV-2 recombinant variants (XD and XE) and Omicron subvariants (BA.1 and BA.2).
Keywords: BA.1 subvariant; BA.2 subvariant; Omicron variant; SARS-CoV-2; allele-specific qRT-PCR; mutation; recombinant variant.
Copyright © 2022 Li, Gao, Chen, Cheng, Niu, Zhang, Yang, Yuan, Xia, Mao, Liu, Dong and Wu.
Conflict of interest statement
Authors GM and HL were employed by Shanxi Guoxin Caregeno Biotechnology Co., Ltd, Taiyuan, China. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Development of multiplex allele-specific RT-qPCR assays for differentiation of SARS-CoV-2 Omicron subvariants.Appl Microbiol Biotechnol. 2024 Dec;108(1):35. doi: 10.1007/s00253-023-12941-2. Epub 2024 Jan 6. Appl Microbiol Biotechnol. 2024. PMID: 38183475
-
Accessible and Adaptable Multiplexed Real-Time PCR Approaches to Identify SARS-CoV-2 Variants of Concern.Microbiol Spectr. 2022 Oct 26;10(5):e0322222. doi: 10.1128/spectrum.03222-22. Epub 2022 Sep 15. Microbiol Spectr. 2022. PMID: 36106882 Free PMC article.
-
[Rapid detection and genotyping of SARS-CoV-2 Omicron BA.4/5 variants using a RT-PCR and CRISPR-Cas12a-based assay].Nan Fang Yi Ke Da Xue Xue Bao. 2023 Apr 20;43(4):516-526. doi: 10.12122/j.issn.1673-4254.2023.04.03. Nan Fang Yi Ke Da Xue Xue Bao. 2023. PMID: 37202186 Free PMC article. Chinese.
-
Current strategies for SARS-CoV-2 molecular detection.Anal Methods. 2022 Nov 24;14(45):4625-4642. doi: 10.1039/d2ay01313d. Anal Methods. 2022. PMID: 36349688 Review.
-
Immunological Studies to Understand Hybrid/Recombinant Variants of SARS-CoV-2.Vaccines (Basel). 2022 Dec 25;11(1):45. doi: 10.3390/vaccines11010045. Vaccines (Basel). 2022. PMID: 36679891 Free PMC article. Review.
Cited by
-
Application of novel dark-quencher labeled probes in multiplex qRT-PCR assays for rapid detection of SARS-CoV-2 variants.J Thorac Dis. 2025 Apr 30;17(4):2159-2173. doi: 10.21037/jtd-24-853. Epub 2025 Apr 28. J Thorac Dis. 2025. PMID: 40400917 Free PMC article.
-
Efficient Tracing of the SARS-CoV-2 Omicron Variants in Santa Barbara County Using a Rapid Quantitative Reverse Transcription PCR Assay.Diagnostics (Basel). 2022 Nov 15;12(11):2805. doi: 10.3390/diagnostics12112805. Diagnostics (Basel). 2022. PMID: 36428863 Free PMC article.
-
Development of multiplex allele-specific RT-qPCR assays for differentiation of SARS-CoV-2 Omicron subvariants.Appl Microbiol Biotechnol. 2024 Dec;108(1):35. doi: 10.1007/s00253-023-12941-2. Epub 2024 Jan 6. Appl Microbiol Biotechnol. 2024. PMID: 38183475
References
-
- Bartsch Y., Cizmeci D., Kang J., Gao H., Shi W., Chandrashekar A. (2022). BA.2 evasion of vaccine induced binding and functional non-neutralizing antibodies. medRxiv. doi: 10.1101/2022.02.25.22271511 - DOI
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

