Compellingly high SARS-CoV-2 susceptibility of Golden Syrian hamsters suggests multiple zoonotic infections of pet hamsters during the COVID-19 pandemic
- PMID: 36064749
- PMCID: PMC9442591
- DOI: 10.1038/s41598-022-19222-4
Compellingly high SARS-CoV-2 susceptibility of Golden Syrian hamsters suggests multiple zoonotic infections of pet hamsters during the COVID-19 pandemic
Abstract
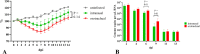
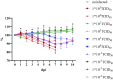
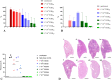
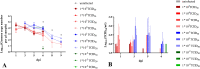
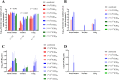
Golden Syrian hamsters (Mesocricetus auratus) are used as a research model for severe acute respiratory syndrome coronavirus type 2 (SARS-CoV-2). Millions of Golden Syrian hamsters are also kept as pets in close contact to humans. To determine the minimum infective dose (MID) for assessing the zoonotic transmission risk, and to define the optimal infection dose for experimental studies, we orotracheally inoculated hamsters with SARS-CoV-2 doses from 1 * 105 to 1 * 10-4 tissue culture infectious dose 50 (TCID50). Body weight and virus shedding were monitored daily. 1 * 10-3 TCID50 was defined as the MID, and this was still sufficient to induce virus shedding at levels up to 102.75 TCID50/ml, equaling the estimated MID for humans. Virological and histological data revealed 1 * 102 TCID50 as the optimal dose for experimental infections. This compelling high susceptibility leading to productive infections in Golden Syrian hamsters must be considered as a potential source of SARS-CoV-2 infection for humans that come into close contact with pet hamsters.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Sex Differences in Lung Imaging and SARS-CoV-2 Antibody Responses in a COVID-19 Golden Syrian Hamster Model.mBio. 2021 Aug 31;12(4):e0097421. doi: 10.1128/mBio.00974-21. Epub 2021 Jul 13. mBio. 2021. PMID: 34253053 Free PMC article.
-
[Specific features of the pathology of the respiratory system in SARS-CoV-2 (Coronaviridae: Coronavirinae: Betacoronavirus: Sarbecovirus) infected Syrian hamsters (Mesocricetus auratus)].Vopr Virusol. 2022 Jan 8;66(6):442-451. doi: 10.36233/0507-4088-63. Vopr Virusol. 2022. PMID: 35019251 Russian.
-
Pathogenesis and transmission of SARS-CoV-2 in golden hamsters.Nature. 2020 Jul;583(7818):834-838. doi: 10.1038/s41586-020-2342-5. Epub 2020 May 14. Nature. 2020. PMID: 32408338 Free PMC article.
-
Hamsters as a Model of Severe Acute Respiratory Syndrome Coronavirus-2.Comp Med. 2021 Oct 1;71(5):398-410. doi: 10.30802/AALAS-CM-21-000036. Epub 2021 Sep 29. Comp Med. 2021. PMID: 34588095 Free PMC article. Review.
-
Animals and SARS-CoV-2: Species susceptibility and viral transmission in experimental and natural conditions, and the potential implications for community transmission.Transbound Emerg Dis. 2021 Jul;68(4):1850-1867. doi: 10.1111/tbed.13885. Epub 2020 Nov 4. Transbound Emerg Dis. 2021. PMID: 33091230 Free PMC article. Review.
Cited by
-
3D imaging of SARS-CoV-2 infected hamster lungs by X-ray phase contrast tomography enables drug testing.Sci Rep. 2024 May 29;14(1):12348. doi: 10.1038/s41598-024-61746-4. Sci Rep. 2024. PMID: 38811688 Free PMC article.
-
SARS-CoV-2 as a Zooanthroponotic Infection: Spillbacks, Secondary Spillovers, and Their Importance.Microorganisms. 2022 Oct 31;10(11):2166. doi: 10.3390/microorganisms10112166. Microorganisms. 2022. PMID: 36363758 Free PMC article. Review.
-
Increased Susceptibility of Rousettus aegyptiacus Bats to Respiratory SARS-CoV-2 Challenge Despite Its Distinct Tropism for Gut Epithelia in Bats.Viruses. 2024 Oct 31;16(11):1717. doi: 10.3390/v16111717. Viruses. 2024. PMID: 39599832 Free PMC article.
-
Rehoming and Other Refinements and Replacement in Procedures Using Golden Hamsters in SARS-CoV-2 Vaccine Research.Animals (Basel). 2023 Aug 14;13(16):2616. doi: 10.3390/ani13162616. Animals (Basel). 2023. PMID: 37627407 Free PMC article.
-
Evidence for SARS-CoV-2 infected Golden Syrian hamsters (Mesocricetus auratus) reducing daily energy expenditure and body core temperature.Sci Rep. 2024 Oct 6;14(1):23263. doi: 10.1038/s41598-024-73765-2. Sci Rep. 2024. PMID: 39370428 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

