KidA, a multi-PAS domain protein, tunes the period of the cyanobacterial circadian oscillator
- PMID: 36067319
- PMCID: PMC9478674
- DOI: 10.1073/pnas.2202426119
KidA, a multi-PAS domain protein, tunes the period of the cyanobacterial circadian oscillator
Abstract
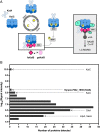
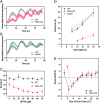
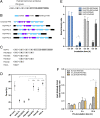
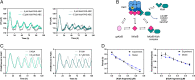
The cyanobacterial clock presents a unique opportunity to understand the biochemical basis of circadian rhythms. The core oscillator, composed of the KaiA, KaiB, and KaiC proteins, has been extensively studied, but a complete picture of its connection to the physiology of the cell is lacking. To identify previously unknown components of the clock, we used KaiB locked in its active fold as bait in an immunoprecipitation/mass spectrometry approach. We found that the most abundant interactor, other than KaiC, was a putative diguanylate cyclase protein predicted to contain multiple Per-Arnt-Sim (PAS) domains, which we propose to name KidA. Here we show that KidA directly binds to the fold-switched active form of KaiB through its N-terminal PAS domains. We found that KidA shortens the period of the circadian clock both in vivo and in vitro and alters the ability of the clock to entrain to light-dark cycles. The dose-dependent effect of KidA on the clock period could be quantitatively recapitulated by a mathematical model in which KidA stabilizes the fold-switched form of KaiB, favoring rebinding to KaiC. Put together, our results show that the period and amplitude of the clock can be modulated by regulating the access of KaiB to the fold-switched form.
Keywords: circadian rhythms; cyanobacteria; diguanylate cyclase.
Conflict of interest statement
The authors declare no competing interest.
Figures
References
-
- Nakajima M., et al. , Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005). - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

