Spike protein-independent attenuation of SARS-CoV-2 Omicron variant in laboratory mice
- PMID: 36075211
- PMCID: PMC9420700
- DOI: 10.1016/j.celrep.2022.111359
Spike protein-independent attenuation of SARS-CoV-2 Omicron variant in laboratory mice
Abstract
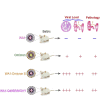
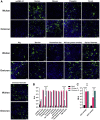
Despite being more transmissible, the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron variant only causes milder diseases in laboratory animals, often accompanied by a lower viral load compared with previous variants of concern. In this study, we report the structural basis for a robust interaction between the receptor-binding domain of the Omicron spike protein and mouse ACE2. We show that pseudovirus bearing the Omicron spike protein efficiently utilizes mouse ACE2 for entry. By comparing viral load and disease severity among laboratory mice infected by a natural Omicron variant or recombinant ancestral viruses bearing either the entire Omicron spike or only the N501Y/Q493R mutations in its spike, we find that mutations outside the spike protein in the Omicron variant may be responsible for the observed lower viral load. Together, our results imply that a post-entry block to the Omicron variant exists in laboratory mice.
Keywords: Balb/c mice; CP: Microbiology; K18-hACE2; Omicron variant; SARS-CoV-2; attenuation.
Published by Elsevier Inc.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



References
-
- Abdelnabi R., Foo C.S., Zhang X., Lemmens V., Maes P., Slechten B., Raymenants J., André E., Weynand B., Dallemier K., Neyts J. The omicron (B.1.1.529) SARS-CoV-2 variant of concern does not readily infect Syrian hamsters. bioRxiv. 2021 doi: 10.1101/2021.12.24.474086. Preprint at. - DOI - PMC - PubMed
-
- Allen M.P., Dominic J. Oxford University Press; 1987. Computer Simulation of Liquids.
-
- Beglov D., Roux B. Finite representation of an infinite bulk system: solvent boundary potential for computer simulations. J. Chem. Phys. 1994;100:9050–9063.
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

