Glycan-Adhering Lectins and Experimental Evaluation of a Lectin FimH Inhibitor in Enterohemorrhagic Escherichia coli (EHEC) O157:H7 Strain EDL933
- PMID: 36077327
- PMCID: PMC9455959
- DOI: 10.3390/ijms23179931
Glycan-Adhering Lectins and Experimental Evaluation of a Lectin FimH Inhibitor in Enterohemorrhagic Escherichia coli (EHEC) O157:H7 Strain EDL933
Abstract
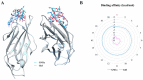
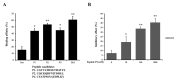
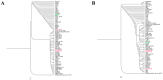
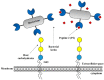
In this study, we tried to develop a FimH inhibitor that inhibits adhesion of enterohemorrhagic Escherichia coli (EHEC) on the epithelium of human intestine during the initial stage of infections. Using a T7 phage display method with a reference strain, EHEC EDL933, FimH was selected as an adherent lectin to GM1a and Gb3 glycans. In order to detect the ligand binding domain (LBD) of FimH, we used a docking simulation and found three binding site sequences of FimH, i.e., P1, P2, and P3. Among Gb3 mimic peptides, P2 was found to have the strongest binding strength. Moreover, in vitro treatment with peptide P2 inhibited binding activity in a concentration-dependent manner. Furthermore, we conducted confirmation experiments through several strains isolated from patients in Korea, EHEC NCCP15736, NCCP15737, and NCCP15739. In addition, we analyzed the evolutionary characteristics of the predicted FimH lectin-like adhesins to construct a lectin-glycan interaction (LGI). We selected 70 recently differentiated strains from the phylogenetic tree of 2240 strains with Shiga toxin in their genome. We can infer EHEC strains dynamically evolved but FimH was conserved during the evolution time according to the phylogenetic tree. Furthermore, FimH could be a reliable candidate of drug target in terms of evolution. We examined how pathogen lectins interact with host glycans early in infection in EDL933 as well as several field strains and confirmed that glycan-like peptides worked as an initial infection inhibitor.
Keywords: FimH; Gb3; enterohemorrhagic Escherichia coli (EHEC); glycan; lectin.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

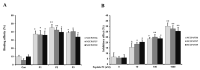
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

