The microbiome-derived metabolite TMAO drives immune activation and boosts responses to immune checkpoint blockade in pancreatic cancer
- PMID: 36083892
- PMCID: PMC9925043
- DOI: 10.1126/sciimmunol.abn0704
The microbiome-derived metabolite TMAO drives immune activation and boosts responses to immune checkpoint blockade in pancreatic cancer
Abstract
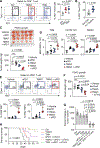
The composition of the gut microbiome can control innate and adaptive immunity and has emerged as a key regulator of tumor growth, especially in the context of immune checkpoint blockade (ICB) therapy. However, the underlying mechanisms for how the microbiome affects tumor growth remain unclear. Pancreatic ductal adenocarcinoma (PDAC) tends to be refractory to therapy, including ICB. Using a nontargeted, liquid chromatography-tandem mass spectrometry-based metabolomic screen, we identified the gut microbe-derived metabolite trimethylamine N-oxide (TMAO), which enhanced antitumor immunity to PDAC. Delivery of TMAO intraperitoneally or via a dietary choline supplement to orthotopic PDAC-bearing mice reduced tumor growth, associated with an immunostimulatory tumor-associated macrophage (TAM) phenotype, and activated effector T cell response in the tumor microenvironment. Mechanistically, TMAO potentiated the type I interferon (IFN) pathway and conferred antitumor effects in a type I IFN-dependent manner. Delivering TMAO-primed macrophages intravenously produced similar antitumor effects. Combining TMAO with ICB (anti-PD1 and/or anti-Tim3) in a mouse model of PDAC significantly reduced tumor burden and improved survival beyond TMAO or ICB alone. Last, the levels of bacteria containing CutC (an enzyme that generates trimethylamine, the TMAO precursor) correlated with long-term survival in patients with PDAC and improved response to anti-PD1 in patients with melanoma. Together, our study identifies the gut microbial metabolite TMAO as a driver of antitumor immunity and lays the groundwork for potential therapeutic strategies targeting TMAO.
Conflict of interest statement
Competing interests
Authors have no conflicts of interest
Figures






Comment in
-
Microbiome-derived metabolites counteract tumor-induced immunosuppression and boost immune checkpoint blockade.Cell Metab. 2022 Dec 6;34(12):1903-1905. doi: 10.1016/j.cmet.2022.11.010. Cell Metab. 2022. PMID: 36476933
References
-
- Schizas D, Charalampakis N, Kole C, Economopoulou P, Koustas E, Gkotsis E, Ziogas D, Psyrri A, Karamouzis MV, Immunotherapy for pancreatic cancer: A 2020 update. Cancer Treat Rev 86, 102016 (2020). - PubMed
-
- Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL, Hwu P, Drake CG, Camacho LH, Kauh J, Odunsi K, Pitot HC, Hamid O, Bhatia S, Martins R, Eaton K, Chen S, Salay TM, Alaparthy S, Grosso JF, Korman AJ, Parker SM, Agrawal S, Goldberg SM, Pardoll DM, Gupta A, Wigginton JM, Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med 366, 2455–2465 (2012). - PMC - PubMed
-
- Hezaveh K, Shinde RS, Klotgen A, Halaby MJ, Lamorte S, Ciudad MT, Quevedo R, Neufeld L, Liu ZQ, Jin R, Grunwald BT, Foerster EG, Chaharlangi D, Guo M, Makhijani P, Zhang X, Pugh TJ, Pinto DM, Co IL, McGuigan AP, Jang GH, Khokha R, Ohashi PS, O’Kane GM, Gallinger S, Navarre WW, Maughan H, Philpott DJ, Brooks DG, McGaha TL, Tryptophan-derived microbial metabolites activate the aryl hydrocarbon receptor in tumor-associated macrophages to suppress anti-tumor immunity. Immunity 55, 324–340 e328 (2022). - PMC - PubMed
-
- Panni RZ, Herndon JM, Zuo C, Hegde S, Hogg GD, Knolhoff BL, Breden MA, Li X, Krisnawan VE, Khan SQ, Schwarz JK, Rogers BE, Fields RC, Hawkins WG, Gupta V, DeNardo DG, Agonism of CD11b reprograms innate immunity to sensitize pancreatic cancer to immunotherapies. Sci Transl Med 11, (2019). - PMC - PubMed
-
- . Li J, Byrne KT, Yan F, Yamazoe T, Chen Z, Baslan T, Richman LP, Lin JH, Sun YH, Rech AJ, Balli D, Hay CA, Sela Y, Merrell AJ, Liudahl SM, Gordon N, Norgard RJ, Yuan S, Yu S, Chao T, Ye S, Eisinger-Mathason TSK, Faryabi RB, Tobias JW, Lowe SW, Coussens LM, Wherry EJ, Vonderheide RH, Stanger BZ, Tumor Cell-Intrinsic Factors Underlie Heterogeneity of Immune Cell Infiltration and Response to Immunotherapy. Immunity 49, 178–193 e177 (2018). - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials