Molecular epidemiology and genetic characterization of SARS-CoV-2 in Kuwait: A descriptive study
- PMID: 36090111
- PMCID: PMC9459148
- DOI: 10.3389/fmicb.2022.858770
Molecular epidemiology and genetic characterization of SARS-CoV-2 in Kuwait: A descriptive study
Abstract
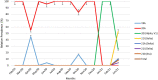
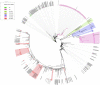
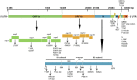
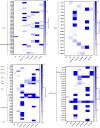
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) has been fatal to human health, affecting almost the entire world. Here we reported, for the first time, characterization of the genetic variants of SARS-CoV-2 circulating in Kuwait to understand their genetic diversity and monitor the accumulation of mutations over time. This study randomly enrolled 209 COVID-19 patients whose nasopharyngeal swabs were positive for SARS-CoV-2 between February 2020 and June 2021 using RT-PCR. The whole genomes of SARS-CoV-2 from the nasopharyngeal swabs were sequenced using the Oxford Nanopore sequencing technology following the ARTIC network protocol. Whole-genome sequencing has identified different clades/sub-clades circulating in Kuwait, mimicking the virus's global spread. Clade 20A was dominant from February 2020 until January 2021, and then clade 20I (Alpha, V1) emerged and dominated. In June 2021, the number of cases infected with clades 21I, 21A, and 21 J (Delta) increased and dominated. We detected several known clade-defining missense and synonymous mutations and other missense mutations in the genes encoding important viral proteins, including ORF1a, S, ORF3a, ORF8 regions and a novel mutation in the N region. ORF1ab region harbored more mutations and deletions (n = 62, 49.2%) compared to the other 12 gene regions, and the most prevalent missense mutations were P314L (97%) in ORF1b and D614G (97%) in the S glycoprotein regions. Detecting and analyzing mutations and monitoring the evolution of SARS-CoV-2 over time is essential to help better understand the spread of various clades/strains of SARS-CoV-2 and their implications for pathogenesis. In addition, knowledge of the circulating variants and genome sequence variability of SARS-CoV-2 may potentially influence the development of vaccines and antiviral drugs to control the COVID-19 pandemic.
Keywords: Kuwait; Nanopore sequencing technology; SARS-CoV-2; molecular epidemiology; variants.
Copyright © 2022 Madi, Safar, Mustafa, Chehadeh, Asadzadeh, Sadeq, Alawadhi, Al-Muhaini and Benthani.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Strain Variation Based on Spike Glycoprotein Gene of SARS-CoV-2 in Kuwait from 2020 to 2021.Pathogens. 2022 Aug 29;11(9):985. doi: 10.3390/pathogens11090985. Pathogens. 2022. PMID: 36145416 Free PMC article.
-
[Genomic Characterization of SARS-CoV-2 Isolates Obtained from Antalya, Türkiye].Mikrobiyol Bul. 2024 Oct;58(4):433-447. doi: 10.5578/mb.20249643. Mikrobiyol Bul. 2024. PMID: 39745215 Turkish.
-
Genome sequence diversity of SARS-CoV-2 in Serbia: insights gained from a 3-year pandemic study.Front Microbiol. 2024 Feb 27;15:1332276. doi: 10.3389/fmicb.2024.1332276. eCollection 2024. Front Microbiol. 2024. PMID: 38476954 Free PMC article.
-
Molecular evolution of SARS-CoV-2 from December 2019 to August 2022.J Med Virol. 2023 Jan;95(1):e28366. doi: 10.1002/jmv.28366. J Med Virol. 2023. PMID: 36458547 Free PMC article. Review.
-
Emergence, evolution, and vaccine production approaches of SARS-CoV-2 virus: Benefits of getting vaccinated and common questions.Saudi J Biol Sci. 2022 Apr;29(4):1981-1997. doi: 10.1016/j.sjbs.2021.12.020. Epub 2021 Dec 13. Saudi J Biol Sci. 2022. PMID: 34924802 Free PMC article. Review.
Cited by
-
Retrospective Cohort Study Comparing the Clinical Profile and Outcomes of Critically Ill Pregnant Patients in Kuwait during the COVID-19 Pandemic Waves.Med Princ Pract. 2024;33(5):441-451. doi: 10.1159/000539004. Epub 2024 Apr 20. Med Princ Pract. 2024. PMID: 38643766 Free PMC article.
-
Comparative Evolutionary Epidemiology of SARS-CoV-2 Delta and Omicron Variants in Kuwait.Viruses. 2024 Nov 30;16(12):1872. doi: 10.3390/v16121872. Viruses. 2024. PMID: 39772182 Free PMC article.
References
-
- Barona-Gómez F., Delaye L., Díaz-Valenzuela E., Plisson F., Cruz-Pérez A., Díaz-Sánchez M., et al. . (2021). Phylogenomics and population genomics of SARS-CoV-2 in Mexico during the pre-vaccination stage reveals variants of interest B.1.1.28.4 and B.1.1.222 or B.1.1.519 and the nucleocapsid mutation S194L associated with symptoms. Microb. Genomics 7:684. doi: 10.1099/mgen.0.000684, PMID: - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

