Pancancer analysis of a potential gene mutation model in the prediction of immunotherapy outcomes
- PMID: 36092890
- PMCID: PMC9459043
- DOI: 10.3389/fgene.2022.917118
Pancancer analysis of a potential gene mutation model in the prediction of immunotherapy outcomes
Abstract
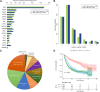
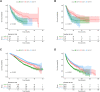
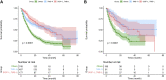
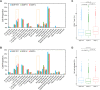
Background: Immune checkpoint blockade (ICB) represents a promising treatment for cancer, but predictive biomarkers are needed. We aimed to develop a cost-effective signature to predict immunotherapy benefits across cancers. Methods: We proposed a study framework to construct the signature. Specifically, we built a multivariate Cox proportional hazards regression model with LASSO using 80% of an ICB-treated cohort (n = 1661) from MSKCC. The desired signature named SIGP was the risk score of the model and was validated in the remaining 20% of patients and an external ICB-treated cohort (n = 249) from DFCI. Results: SIGP was based on 18 candidate genes (NOTCH3, CREBBP, RNF43, PTPRD, FAM46C, SETD2, PTPRT, TERT, TET1, ROS1, NTRK3, PAK7, BRAF, LATS1, IL7R, VHL, TP53, and STK11), and we classified patients into SIGP high (SIGP-H), SIGP low (SIGP-L) and SIGP wild type (SIGP-WT) groups according to the SIGP score. A multicohort validation demonstrated that patients in SIGP-L had significantly longer overall survival (OS) in the context of ICB therapy than those in SIGP-WT and SIGP-H (44.00 months versus 13.00 months and 14.00 months, p < 0.001 in the test set). The survival of patients grouped by SIGP in non-ICB-treated cohorts was different, and SIGP-WT performed better than the other groups. In addition, SIGP-L + TMB-L (approximately 15% of patients) had similar survivals to TMB-H, and patients with both SIGP-L and TMB-H had better survival. Further analysis on tumor-infiltrating lymphocytes demonstrated that the SIGP-L group had significantly increased abundances of CD8+ T cells. Conclusion: Our proposed model of the SIGP signature based on 18-gene mutations has good predictive value for the clinical benefit of ICB in pancancer patients. Additional patients without TMB-H were identified by SIGP as potential candidates for ICB, and the combination of both signatures showed better performance than the single signature.
Keywords: biomarker; gene mutation signature; immune checkpoint blockade; immunotherapy; statistical modeling.
Copyright © 2022 Yu and Gong.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
References
-
- Algamal Z. Y., Lee M. H. (2015). Penalized logistic regression with the adaptive LASSO for gene selection in high-dimensional cancer classification. Expert Syst. Appl. 42 (23), 9326–9332. 10.1016/j.eswa.2015.08.016 - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

