Structure and Function of the Nuclear Pore Complex
- PMID: 36096637
- PMCID: PMC9732903
- DOI: 10.1101/cshperspect.a041264
Structure and Function of the Nuclear Pore Complex
Abstract
The nucleus, a genome-containing organelle eponymous of eukaryotes, is enclosed by a double membrane continuous with the endoplasmic reticulum. The nuclear pore complex (NPC) is an ∼110-MDa, ∼1000-protein channel that selectively transports macromolecules across the nuclear envelope and thus plays a central role in the regulated flow of genetic information from transcription to translation. Its size, complexity, and flexibility have hindered determination of atomistic structures of intact NPCs. Recent studies have overcome these hurdles by combining biochemical reconstitution and docking of high-resolution structures of NPC subcomplexes into cryo-electron tomographic reconstructions with biochemical and physiological validation. Here, we provide an overview of the near-atomic composite structure of the human NPC, a milestone toward unlocking a molecular understanding of mRNA export, NPC-associated diseases, and viral host-pathogen interactions, serving as a paradigm for studying similarly large complexes.
Copyright © 2022 Cold Spring Harbor Laboratory Press; all rights reserved.
Figures
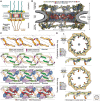




Similar articles
-
Architecture of the linker-scaffold in the nuclear pore.Science. 2022 Jun 10;376(6598):eabm9798. doi: 10.1126/science.abm9798. Epub 2022 Jun 10. Science. 2022. PMID: 35679425 Free PMC article.
-
The Structure of the Nuclear Pore Complex (An Update).Annu Rev Biochem. 2019 Jun 20;88:725-783. doi: 10.1146/annurev-biochem-062917-011901. Epub 2019 Mar 18. Annu Rev Biochem. 2019. PMID: 30883195 Free PMC article. Review.
-
AI-based structure prediction empowers integrative structural analysis of human nuclear pores.Science. 2022 Jun 10;376(6598):eabm9506. doi: 10.1126/science.abm9506. Epub 2022 Jun 10. Science. 2022. PMID: 35679397
-
Structural analysis of the nuclear pore complex by integrated approaches.Curr Opin Struct Biol. 2009 Apr;19(2):226-32. doi: 10.1016/j.sbi.2009.02.009. Epub 2009 Mar 25. Curr Opin Struct Biol. 2009. PMID: 19327984 Review.
-
Nuclear pore complex: biochemistry and biophysics of nucleocytoplasmic transport in health and disease.Int Rev Cell Mol Biol. 2011;287:233-86. doi: 10.1016/B978-0-12-386043-9.00006-2. Int Rev Cell Mol Biol. 2011. PMID: 21414590 Review.
Cited by
-
Structural basis for a nucleoporin exportin complex between RanBP2, SUMO1-RanGAP1, the E2 Ubc9, Crm1 and the Ran GTPase.bioRxiv [Preprint]. 2024 Dec 26:2024.10.04.616749. doi: 10.1101/2024.10.04.616749. bioRxiv. 2024. Update in: Nat Commun. 2025 Jul 11;16(1):6403. doi: 10.1038/s41467-025-61694-1. PMID: 39763778 Free PMC article. Updated. Preprint.
-
Mechanisms of PP2A-Ankle2 dependent nuclear reassembly after mitosis.Elife. 2025 Feb 18;13:RP104233. doi: 10.7554/eLife.104233. Elife. 2025. PMID: 39964262 Free PMC article.
-
Particle fusion of super-resolution data reveals the unit structure of Nup96 in Nuclear Pore Complex.Sci Rep. 2023 Aug 16;13(1):13327. doi: 10.1038/s41598-023-39829-5. Sci Rep. 2023. PMID: 37587192 Free PMC article.
-
Nuclear basket proteins regulate the distribution and mobility of nuclear pore complexes in budding yeast.Mol Biol Cell. 2024 Nov 1;35(11):ar143. doi: 10.1091/mbc.E24-08-0371. Epub 2024 Sep 25. Mol Biol Cell. 2024. PMID: 39320946 Free PMC article.
-
Strategies for the Viral Exploitation of Nuclear Pore Transport Pathways.Viruses. 2025 Jan 23;17(2):151. doi: 10.3390/v17020151. Viruses. 2025. PMID: 40006906 Free PMC article. Review.
References
-
- Akey CW. 1995. Structural plasticity of the nuclear pore complex. J Mol Biol 248: 273–293. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
