The choanoflagellate pore-forming lectin SaroL-1 punches holes in cancer cells by targeting the tumor-related glycosphingolipid Gb3
- PMID: 36097056
- PMCID: PMC9468336
- DOI: 10.1038/s42003-022-03869-w
The choanoflagellate pore-forming lectin SaroL-1 punches holes in cancer cells by targeting the tumor-related glycosphingolipid Gb3
Abstract
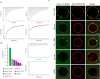
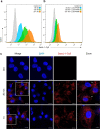
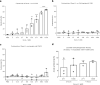
Choanoflagellates are primitive protozoa used as models for animal evolution. They express a large variety of multi-domain proteins contributing to adhesion and cell communication, thereby providing a rich repertoire of molecules for biotechnology. Adhesion often involves proteins adopting a β-trefoil fold with carbohydrate-binding properties therefore classified as lectins. Sequence database screening with a dedicated method resulted in TrefLec, a database of 44714 β-trefoil candidate lectins across 4497 species. TrefLec was searched for original domain combinations, which led to single out SaroL-1 in the choanoflagellate Salpingoeca rosetta, that contains both β-trefoil and aerolysin-like pore-forming domains. Recombinant SaroL-1 is shown to bind galactose and derivatives, with a stronger affinity for cancer-related α-galactosylated epitopes such as the glycosphingolipid Gb3, when embedded in giant unilamellar vesicles or cell membranes. Crystal structures of complexes with Gb3 trisaccharide and GalNAc provided the basis for building a model of the oligomeric pore. Finally, recognition of the αGal epitope on glycolipids required for hemolysis of rabbit erythrocytes suggests that toxicity on cancer cells is achieved through carbohydrate-dependent pore-formation.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


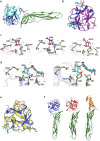


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

