Impaired XK recycling for importing manganese underlies striatal vulnerability in Huntington's disease
- PMID: 36099524
- PMCID: PMC9475296
- DOI: 10.1083/jcb.202112073
Impaired XK recycling for importing manganese underlies striatal vulnerability in Huntington's disease
Abstract
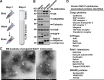
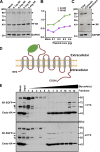
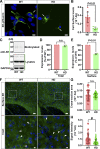
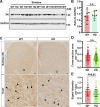
Mutant huntingtin, which causes Huntington's disease (HD), is ubiquitously expressed but induces preferential loss of striatal neurons by unclear mechanisms. Rab11 dysfunction mediates homeostatic disturbance of HD neurons. Here, we report that Rab11 dysfunction also underscores the striatal vulnerability in HD. We profiled the proteome of Rab11-positive endosomes of HD-vulnerable striatal cells to look for protein(s) linking Rab11 dysfunction to striatal vulnerability in HD and found XK, which triggers the selective death of striatal neurons in McLeod syndrome. XK was trafficked together with Rab11 and was diminished on the surface of immortalized HD striatal cells and striatal neurons in HD mouse brains. We found that XK participated in transporting manganese, an essential trace metal depleted in HD brains. Introducing dominantly active Rab11 into HD striatal cells improved XK dynamics and increased manganese accumulation in an XK-dependent manner. Our study suggests that impaired Rab11-based recycling of XK onto cell surfaces for importing manganese is a driver of striatal dysfunction in Huntington's disease.
© 2022 Chhetri et al.
Figures






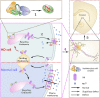
Comment in
-
Endosomal recycling defects link Huntington's disease with McLeod syndrome.J Cell Biol. 2022 Oct 3;221(10):e202208164. doi: 10.1083/jcb.202208164. Epub 2022 Sep 16. J Cell Biol. 2022. PMID: 36112418 Free PMC article.
References
-
- MacDonald M.E., Ambrose C.M., Duyao M.P., Myers R.H., Lin C., Srinidhi L., Barnes G., Taylor S.A., James M., Groot N., et al. 1993. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington’s disease collaborative research group. Cell. 72:971-983. 10.1016/0092-8674(93)90585-e - DOI - PubMed
-
- Bogershausen, N., Shahrzad N., Chong J.X., von Kleist-Retzow J.C., Stanga D., Li Y., Bernier F.P., Loucks C.M., Wirth R., Puffenberger E.G., et al. 2013. Recessive TRAPPC11 mutations cause a disease spectrum of limb girdle muscular dystrophy and myopathy with movement disorder and intellectual disability. Am. J. Hum. Genet. 93:181–190. 10.1016/j.ajhg.2013.05.028 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

