Engineering enhanced thermostability into the Geobacillus pallidus nitrile hydratase
- PMID: 36106339
- PMCID: PMC9465369
- DOI: 10.1016/j.crstbi.2022.07.002
Engineering enhanced thermostability into the Geobacillus pallidus nitrile hydratase
Abstract
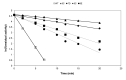
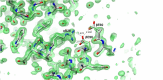
Nitrile hydratases (NHases) are important biocatalysts for the enzymatic conversion of nitriles to industrially-important amides such as acrylamide and nicotinamide. Although thermostability in this enzyme class is generally low, there is not sufficient understanding of its basis for rational enzyme design. The gene expressing the Co-type NHase from the moderate thermophile, Geobacillus pallidus RAPc8 (NRRL B-59396), was subjected to random mutagenesis. Four mutants were selected that were 3 to 15-fold more thermostable than the wild-type NHase, resulting in a 3.4-7.6 kJ/mol increase in the activation energy of thermal inactivation at 63 °C. High resolution X-ray crystal structures (1.15-1.80 Å) were obtained of the wild-type and four mutant enzymes. Mutant 9E, with a resolution of 1.15 Å, is the highest resolution crystal structure obtained for a nitrile hydratase to date. Structural comparisons between the wild-type and mutant enzymes illustrated the importance of salt bridges and hydrogen bonds in enhancing NHase thermostability. These additional interactions variously improved thermostability by increased intra- and inter-subunit interactions, preventing cooperative unfolding of α-helices and stabilising loop regions. Some hydrogen bonds were mediated via a water molecule, specifically highlighting the significance of structured water molecules in protein thermostability. Although knowledge of the mutant structures makes it possible to rationalize their behaviour, it would have been challenging to predict in advance that these mutants would be stabilising.
Keywords: Crystal structure; Directed evolution; Electrostatic interactions; Nitrile hydratase; Protein engineering; Protein stability; Random mutagenesis; Thermophile; Thermostability.
© 2022 Published by Elsevier B.V.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures












References
-
- Arnott M.A., Michael R.A., Thompson C.R., Hough D.W., Danson M.J. Thermostability and thermoactivity of citrate synthases from the thermophilic and hyperthermophilic archaea, Thermoplasma acidophilum and Pyrococcus furiosus. J. Mol. Biol. 2000;304:657–668. - PubMed
-
- Bloom J.D., Meyer M.M., Meinhold P., Otey C.R., MacMillan D., Arnold F.H. Evolving strategies for enzyme engineering. Curr. Opin. Struct. Biol. 2005;15:447–452. - PubMed
-
- Bogin O., Peretz M., Hacham Y., Burstein Y., Korkhin Y., Frolow F. Enhanced thermal stability of Clostridium beijerinckii alcohol dehydrogenase after strategic substitution of amino acid residues with prolines from the homologous thermophilic Thermoanaerobacter brockii alcohol dehydrogenase. Protein Sci. 1998;7:1156–1163. - PMC - PubMed
-
- Cameron R.A. University College London (University of London; 2003. Nitrile Degrading Enzymes from Extreme Environments.
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

