Genetic engineering for enhanced production of a novel alkaline protease BSP-1 in Bacillus amyloliquefaciens
- PMID: 36110310
- PMCID: PMC9468883
- DOI: 10.3389/fbioe.2022.977215
Genetic engineering for enhanced production of a novel alkaline protease BSP-1 in Bacillus amyloliquefaciens
Abstract
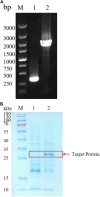
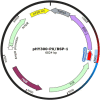
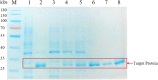
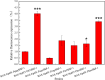
Alkaline protease has been widely applied in food, medicine, environmental protection and other industrial fields. However, the current activity and yield of alkaline protease cannot meet the demand. Therefore, it is important to identify new alkaline proteases with high activity. In this study, we cloned a potential alkaline protease gene bsp-1 from a Bacillus subtilis strain isolated in our laboratory. BSP-1 shows the highest sequence similarity to subtilisin NAT (S51909) from B. subtilis natto. Then, we expressed BSP-1 in Bacillus amyloliquefaciens BAX-9 and analyzed the protein expression level under a collection of promoters. The results show that the P43 promoter resulted in the highest transcription level, protein level and enzyme activity. Finally, we obtained a maximum activity of 524.12 U/mL using the P43 promoter after fermentation medium optimization. In conclusion, this study identified an alkaline protease gene bsp-1 from B. subtilis and provided a new method for high-efficiency alkaline protease expression in B. amyloliquefaciens.
Keywords: alkaline protease; bacillus amyloliquefaciens; fermentation optimization; promoter screening; recombinant expression.
Copyright © 2022 Jiang, Ye, Liu, Huang, Jiang, Zou, Li, Han and Wei.
Conflict of interest statement
YL was employed by GeneMind Biosciences Company Limited. CY and KH were employed by China National Tobacco Corporation. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Bacillus amyloliquefaciens: Harnessing Its Potential for Industrial, Medical, and Agricultural Applications-A Comprehensive Review.Microorganisms. 2023 Aug 31;11(9):2215. doi: 10.3390/microorganisms11092215. Microorganisms. 2023. PMID: 37764059 Free PMC article. Review.
-
Engineering of a Bacillus amyloliquefaciens Strain with High Neutral Protease Producing Capacity and Optimization of Its Fermentation Conditions.PLoS One. 2016 Jan 11;11(1):e0146373. doi: 10.1371/journal.pone.0146373. eCollection 2016. PLoS One. 2016. PMID: 26752595 Free PMC article.
-
Homology modeling and heterologous expression of highly alkaline subtilisin-like serine protease from Bacillus halodurans C-125.Biotechnol Lett. 2021 Feb;43(2):479-494. doi: 10.1007/s10529-020-03025-6. Epub 2020 Oct 12. Biotechnol Lett. 2021. PMID: 33047274
-
Development of Bacillus amyloliquefaciens as a high-level recombinant protein expression system.J Ind Microbiol Biotechnol. 2019 Jan;46(1):113-123. doi: 10.1007/s10295-018-2089-2. Epub 2018 Nov 7. J Ind Microbiol Biotechnol. 2019. PMID: 30406346
-
Microbial alkaline proteases: Optimization of production parameters and their properties.J Genet Eng Biotechnol. 2017 Jun;15(1):115-126. doi: 10.1016/j.jgeb.2017.02.001. Epub 2017 Feb 27. J Genet Eng Biotechnol. 2017. PMID: 30647648 Free PMC article. Review.
Cited by
-
Screening of Bacillus spp. bacterial endophytes for protease production, and application in feather degradation and bio-detergent additive.Heliyon. 2024 May 4;10(9):e30736. doi: 10.1016/j.heliyon.2024.e30736. eCollection 2024 May 15. Heliyon. 2024. PMID: 38765083 Free PMC article.
-
Bacillus amyloliquefaciens: Harnessing Its Potential for Industrial, Medical, and Agricultural Applications-A Comprehensive Review.Microorganisms. 2023 Aug 31;11(9):2215. doi: 10.3390/microorganisms11092215. Microorganisms. 2023. PMID: 37764059 Free PMC article. Review.
-
Genetic identification and expression optimization of a novel protease HapR from Bacillus velezensis.Front Bioeng Biotechnol. 2024 Mar 13;12:1383083. doi: 10.3389/fbioe.2024.1383083. eCollection 2024. Front Bioeng Biotechnol. 2024. PMID: 38544979 Free PMC article.
References
-
- Agrebi R., Haddar A., Hmidet N., Jellouli K., Manni L., Nasri M. (2009). BSF1 fibrinolytic enzyme from a marine bacterium Bacillus subtilis A26: Purification, biochemical and molecular characterization. Process Biochem. 44 (11), 1252–1259. 10.1016/j.procbio.2009.06.024 - DOI
LinkOut - more resources
Full Text Sources
Research Materials

