Therapeutic and prognostic potential of GPCRs in prostate cancer from multi-omics landscape
- PMID: 36110544
- PMCID: PMC9468875
- DOI: 10.3389/fphar.2022.997664
Therapeutic and prognostic potential of GPCRs in prostate cancer from multi-omics landscape
Abstract
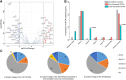
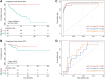
Prostate cancer (PRAD) is a common and fatal malignancy. It is difficult to manage clinically due to drug resistance and poor prognosis, thus creating an urgent need for novel therapeutic targets and prognostic biomarkers. Although G protein-coupled receptors (GPCRs) have been most attractive for drug development, there have been lack of an exhaustive assessment on GPCRs in PRAD like their molecular features, prognostic and therapeutic values. To close this gap, we herein systematically investigate multi-omics profiling for GPCRs in the primary PRAD by analyzing somatic mutations, somatic copy-number alterations (SCNAs), DNA methylation and mRNA expression. GPCRs exhibit low expression levels and mutation frequencies while SCNAs are more prevalent. 46 and 255 disease-related GPCRs are identified by the mRNA expression and DNA methylation analysis, respectively, complementing information lack in the genome analysis. In addition, the genomic alterations do not exhibit an observable correlation with the GPCR expression, reflecting the complex regulatory processes from DNA to RNA. Conversely, a tight association is observed between the DNA methylation and mRNA expression. The virtual screening and molecular dynamics simulation further identify four potential drugs in repositioning to PRAD. The combination of 3 clinical characteristics and 26 GPCR molecular features revealed by the transcriptome and genome exhibit good performance in predicting progression-free survival in patients with the primary PRAD, providing candidates as new biomarkers. These observations from the multi-omics analysis on GPCRs provide new insights into the underlying mechanism of primary PRAD and potential of GPCRs in developing therapeutic strategies on PRAD.
Keywords: G protein-coupled receptors; multi-omics; prognostic model; prostate cancer; virtual screening.
Copyright © 2022 Li, Chen, Chen, Yu, Guo, Li and Pu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Multi-omics integration analysis of GPCRs in pan-cancer to uncover inter-omics relationships and potential driver genes.Comput Biol Med. 2023 Jul;161:106988. doi: 10.1016/j.compbiomed.2023.106988. Epub 2023 May 11. Comput Biol Med. 2023. PMID: 37201441
-
Immune infiltration phenotypes of prostate adenocarcinoma and their clinical implications.Cancer Med. 2021 Aug;10(15):5358-5374. doi: 10.1002/cam4.4063. Epub 2021 Jun 15. Cancer Med. 2021. PMID: 34128342 Free PMC article.
-
Identification of Prognostic Biomarkers Associated with Cancer Stem Cell Features in Prostate Adenocarcinoma.Med Sci Monit. 2020 Jul 31;26:e924543. doi: 10.12659/MSM.924543. Med Sci Monit. 2020. PMID: 32735556 Free PMC article.
-
GPCRs in pancreatic adenocarcinoma: Contributors to tumour biology and novel therapeutic targets.Br J Pharmacol. 2020 Jun;177(11):2434-2455. doi: 10.1111/bph.15028. Epub 2020 Apr 12. Br J Pharmacol. 2020. PMID: 32060895 Free PMC article. Review.
-
Accelerating GPCR Drug Discovery With Conformation-Stabilizing VHHs.Front Mol Biosci. 2022 May 23;9:863099. doi: 10.3389/fmolb.2022.863099. eCollection 2022. Front Mol Biosci. 2022. PMID: 35677880 Free PMC article. Review.
Cited by
-
Pharmacological Efficacy of Repurposing Drugs in the Treatment of Prostate Cancer.Int J Mol Sci. 2023 Feb 19;24(4):4154. doi: 10.3390/ijms24044154. Int J Mol Sci. 2023. PMID: 36835564 Free PMC article. Review.
-
Advancements in MRI-Based Radiomics and Artificial Intelligence for Prostate Cancer: A Comprehensive Review and Future Prospects.Cancers (Basel). 2023 Jul 28;15(15):3839. doi: 10.3390/cancers15153839. Cancers (Basel). 2023. PMID: 37568655 Free PMC article. Review.
References
-
- Albanito L., Lappano R., Madeo A., Chimento A., Prossnitz Eric R., Cappello Anna R., et al. (2008). Retracted: G-protein–coupled receptor 30 and estrogen receptor-α are involved in the proliferative effects induced by atrazine in ovarian cancer cells. Environ. Health Perspect. 116 (12), 1648–1655. 10.1289/ehp.11297 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

