Integrating Text Mining into the Curation of Disease Maps
- PMID: 36139119
- PMCID: PMC9496510
- DOI: 10.3390/biom12091278
Integrating Text Mining into the Curation of Disease Maps
Abstract
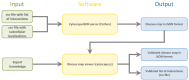
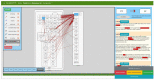
An adequate visualization form is required to gain an overview and ultimately understand the complex and diverse biological mechanisms of diseases. Recently, disease maps have been introduced for this purpose. A disease map is defined as a systems biological map or model that combines metabolic, signaling, and physiological pathways to create a comprehensive overview of known disease mechanisms. With the increase in publications describing biological interactions, efforts in creating and curating comprehensive disease maps is growing accordingly. Therefore, new computational approaches are needed to reduce the time that manual curation takes. Test mining algorithms can be used to analyse the natural language of scientific publications. These types of algorithms can take humanly readable text passages and convert them into a more ordered, machine-usable data structure. To support the creation of disease maps by text mining, we developed an interactive, user-friendly disease map viewer. The disease map viewer displays text mining results in a systems biology map, where the user can review them and either validate or reject identified interactions. Ultimately, the viewer brings together the time-saving advantages of text mining with the accuracy of manual data curation.
Keywords: disease maps; systems biology; text mining.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Mazein A., Ostaszewski M., Kuperstein I., Watterson S., Le Novère N., Lefaudeux D., De Meulder B., Pellet J., Balaur I., Saqi M., et al. Systems medicine disease maps: Community-driven comprehensive representation of disease mechanisms. NPJ Syst. Biol. Appl. 2018;4:21. doi: 10.1038/s41540-018-0059-y. - DOI - PMC - PubMed
-
- Hucka M., Finney A., Sauro H.M., Bolouri H., Doyle J.C., Kitano H., Arkin A.P., Bornstein B.J., Bray D., Cornish-Bowden A., et al. The systems biology markup language (SBML): A medium for representation and exchange of biochemical network models. Bioinformatics. 2003;19:524–531. doi: 10.1093/bioinformatics/btg015. - DOI - PubMed
-
- Ostaszewski M., Niarakis A., Mazein A., Kuperstein I., Phair R., Orta-Resendiz A., Singh V., Aghamiri S.S., Acencio M.L., Glaab E., et al. COVID-19 Disease Map, a computational knowledge repository of virus–host interaction mechanisms. Mol. Syst. Biol. 2021;17 doi: 10.15252/msb.202110851. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

