A Method for Isolation Bacteriophage Particles-Free Genomic DNA, Exemplified by TP-84, Infecting Thermophilic Geobacillus
- PMID: 36144384
- PMCID: PMC9502220
- DOI: 10.3390/microorganisms10091782
A Method for Isolation Bacteriophage Particles-Free Genomic DNA, Exemplified by TP-84, Infecting Thermophilic Geobacillus
Abstract
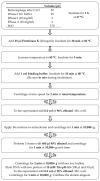
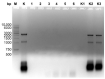
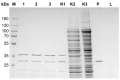
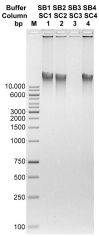
DNA purification methods are indispensable tools of molecular biology, used for many decades. Nevertheless, for certain specialized applications, the currently employed techniques are not sufficiently effective. While examining a number of the existing methods to purify the genomic DNA of the thermophilic bacteriophage TP-84, which infects Geobacillus stearothermophilus (G. stearothermophilus), we have found out that the obtained DNA is contaminated with trace amounts of infectious TP-84 particles. This was detrimental for the bacteriophage genetic manipulation purposes, as finding the recombinant TP-84 clones was essentially impossible due to the appearance of a high background of native bacteriophage plaques. Thus, we have developed a method, which enables the fast and efficient isolation of a bacteriophage genomic DNA from concentrated phage preparations, obtained using CsCl gradient ultracentrifugation, without the need to remove concentrated CsCl solutions. The method employs silica columns and mini-scale isolation of microgram amounts of high quality DNA. It is universal-the silica mini-columns from various manufacturers can be used to conduct the procedure. The purified DNA, free from infectious bacteriophage particles, is ready for further manipulations. This is particularly important for such thermophilic bacteriophages that may partially survive standard isolation procedures and contaminate the final DNA product.
Keywords: DNA purification; TP-84; bacteriophage; genomics; phage display; thermophile.
Conflict of interest statement
The authors declare that there is no conflict of interest regarding the publication of this article. The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

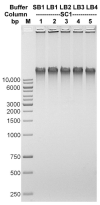
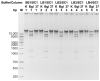

Similar articles
-
A novel thermostable TP-84 capsule depolymerase: a method for rapid polyethyleneimine processing of a bacteriophage-expressed proteins.Microb Cell Fact. 2023 Apr 25;22(1):80. doi: 10.1186/s12934-023-02086-2. Microb Cell Fact. 2023. PMID: 37098567 Free PMC article.
-
A Method for Rapid Polyethyleneimine-Based Purification of Bacteriophage-Expressed Proteins from Diluted Crude Lysates, Exemplified by Thermostable TP-84 Depolymerase.Microorganisms. 2023 Sep 19;11(9):2340. doi: 10.3390/microorganisms11092340. Microorganisms. 2023. PMID: 37764184 Free PMC article.
-
Sequence, genome organization, annotation and proteomics of the thermophilic, 47.7-kb Geobacillus stearothermophilus bacteriophage TP-84 and its classification in the new Tp84virus genus.PLoS One. 2018 Apr 6;13(4):e0195449. doi: 10.1371/journal.pone.0195449. eCollection 2018. PLoS One. 2018. PMID: 29624616 Free PMC article.
-
Bacteriophages of Thermophilic 'Bacillus Group' Bacteria-A Review.Microorganisms. 2021 Jul 16;9(7):1522. doi: 10.3390/microorganisms9071522. Microorganisms. 2021. PMID: 34361957 Free PMC article. Review.
-
Bacteriophages of Thermophilic 'Bacillus Group' Bacteria-A Systematic Review, 2023 Update.Int J Mol Sci. 2024 Mar 8;25(6):3125. doi: 10.3390/ijms25063125. Int J Mol Sci. 2024. PMID: 38542099 Free PMC article.
Cited by
-
Discovery of phage CSF, a novel generalist bacteriophage targeting multidrug-resistant and potentially pathogenic Staphylococcus spp. and Mammaliicoccus spp.Arch Virol. 2025 Jul 29;170(8):188. doi: 10.1007/s00705-025-06370-x. Arch Virol. 2025. PMID: 40731165
-
Advancements in ovarian cancer immunodiagnostics and therapeutics via phage display technology.Front Immunol. 2024 May 28;15:1402862. doi: 10.3389/fimmu.2024.1402862. eCollection 2024. Front Immunol. 2024. PMID: 38863706 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources

