Functional Characterization of an Arabidopsis Profilin Protein as a Molecular Chaperone under Heat Shock Stress
- PMID: 36144503
- PMCID: PMC9504416
- DOI: 10.3390/molecules27185771
Functional Characterization of an Arabidopsis Profilin Protein as a Molecular Chaperone under Heat Shock Stress
Abstract
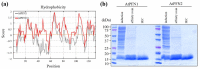
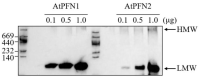
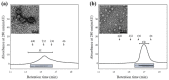
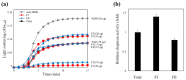
Profilins (PFNs) are actin monomer-binding proteins that function as antimicrobial agents in plant phloem sap. Although the roles of Arabidopsis thaliana profilin protein isoforms (AtPFNs) in regulating actin polymerization have already been described, their biochemical and molecular functions remain to be elucidated. Interestingly, a previous study indicated that AtPFN2 with high molecular weight (HMW) complexes showed lower antifungal activity than AtPFN1 with low molecular weight (LMW). These were bacterially expressed and purified to characterize the unknown functions of AtPFNs with different structures. In this study, we found that AtPFN1 and AtPFN2 proteins have LMW and HMW structures, respectively, but only AtPFN2 has a potential function as a molecular chaperone, which has never been reported elsewhere. AtPFN2 has better protein stability than AtPFN1 due to its higher molecular weight under heat shock conditions. The function of AtPFN2 as a holdase chaperone predominated in the HMW complexes, whereas the chaperone function of AtPFN1 was not observed in the LMW forms. These results suggest that AtPFN2 plays a critical role in plant tolerance by increasing hydrophobicity due to external heat stress.
Keywords: AtPFN; heat shock; higher molecular weight; molecular chaperone; profilin.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





Similar articles
-
Molecular mechanism of Arabidopsis thaliana profilins as antifungal proteins.Biochim Biophys Acta Gen Subj. 2018 Dec;1862(12):2545-2554. doi: 10.1016/j.bbagen.2018.07.028. Epub 2018 Jul 26. Biochim Biophys Acta Gen Subj. 2018. PMID: 30056100
-
Thioredoxin reductase type C (NTRC) orchestrates enhanced thermotolerance to Arabidopsis by its redox-dependent holdase chaperone function.Mol Plant. 2013 Mar;6(2):323-36. doi: 10.1093/mp/sss105. Epub 2012 Sep 30. Mol Plant. 2013. PMID: 23024205
-
Functional changes of OsTrxm from reductase to molecular chaperone under heat shock stress.Plant Physiol Biochem. 2023 Oct;203:108005. doi: 10.1016/j.plaphy.2023.108005. Epub 2023 Sep 7. Plant Physiol Biochem. 2023. PMID: 37776672
-
[Profilins in plant cells].Zhi Wu Sheng Li Yu Fen Zi Sheng Wu Xue Xue Bao. 2006 Jun;32(3):261-70. Zhi Wu Sheng Li Yu Fen Zi Sheng Wu Xue Xue Bao. 2006. PMID: 16775392 Review. Chinese.
-
Actin and actin-binding proteins in higher plants.Protoplasma. 2001;215(1-4):89-104. doi: 10.1007/BF01280306. Protoplasma. 2001. PMID: 11732068 Review.
Cited by
-
Redox dynamics in seeds of Acer spp: unraveling adaptation strategies of different seed categories.Front Plant Sci. 2024 Jul 24;15:1430695. doi: 10.3389/fpls.2024.1430695. eCollection 2024. Front Plant Sci. 2024. PMID: 39114470 Free PMC article.
-
Antifungal Action of Arabidopsis thaliana TCP21 via Induction of Oxidative Stress and Apoptosis.Antioxidants (Basel). 2023 Sep 15;12(9):1767. doi: 10.3390/antiox12091767. Antioxidants (Basel). 2023. PMID: 37760070 Free PMC article.
-
Evolutionary adaptation under climate change: Aedes sp. demonstrates potential to adapt to warming.Proc Natl Acad Sci U S A. 2025 Jan 14;122(2):e2418199122. doi: 10.1073/pnas.2418199122. Epub 2025 Jan 7. Proc Natl Acad Sci U S A. 2025. PMID: 39772738 Free PMC article.
-
Evolutionary adaptation under climate change: Aedes sp. demonstrates potential to adapt to warming.bioRxiv [Preprint]. 2024 Sep 6:2024.08.23.609454. doi: 10.1101/2024.08.23.609454. bioRxiv. 2024. Update in: Proc Natl Acad Sci U S A. 2025 Jan 14;122(2):e2418199122. doi: 10.1073/pnas.2418199122. PMID: 39229052 Free PMC article. Updated. Preprint.
-
The Roadmap of Plant Antimicrobial Peptides Under Environmental Stress: From Farm to Bedside.Probiotics Antimicrob Proteins. 2024 Dec;16(6):2269-2304. doi: 10.1007/s12602-024-10354-9. Epub 2024 Sep 3. Probiotics Antimicrob Proteins. 2024. PMID: 39225894 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

